+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13102 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human LonP1, R-state, incubated in AMPPCP | |||||||||
 Map data Map data | LonP1, R-state, ADP bound, incubated in AMPPCP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | protease / chaperone / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationoxidation-dependent protein catabolic process / response to aluminum ion / PH domain binding / endopeptidase La / mitochondrial protein catabolic process / G-quadruplex DNA binding / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / mitochondrial nucleoid / insulin receptor substrate binding ...oxidation-dependent protein catabolic process / response to aluminum ion / PH domain binding / endopeptidase La / mitochondrial protein catabolic process / G-quadruplex DNA binding / ATP-dependent peptidase activity / protein quality control for misfolded or incompletely synthesized proteins / mitochondrial nucleoid / insulin receptor substrate binding / Mitochondrial unfolded protein response (UPRmt) / chaperone-mediated protein complex assembly / DNA polymerase binding / response to hormone / negative regulation of insulin receptor signaling pathway / Mitochondrial protein degradation / : / mitochondrion organization / ADP binding / single-stranded DNA binding / cellular response to oxidative stress / sequence-specific DNA binding / response to hypoxia / single-stranded RNA binding / mitochondrial matrix / serine-type endopeptidase activity / ATP hydrolysis activity / mitochondrion / nucleoplasm / ATP binding / membrane / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Abrahams JP / Mohammed I / Schmitz KA / Schenck N / Maier T | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Catalytic cycling of human mitochondrial Lon protease. Authors: Inayathulla Mohammed / Kai A Schmitz / Niko Schenck / Dimitrios Balasopoulos / Annika Topitsch / Timm Maier / Jan Pieter Abrahams /  Abstract: The mitochondrial Lon protease (LonP1) regulates mitochondrial health by removing redundant proteins from the mitochondrial matrix. We determined LonP1 in eight nucleotide-dependent conformational ...The mitochondrial Lon protease (LonP1) regulates mitochondrial health by removing redundant proteins from the mitochondrial matrix. We determined LonP1 in eight nucleotide-dependent conformational states by cryoelectron microscopy (cryo-EM). The flexible assembly of N-terminal domains had 3-fold symmetry, and its orientation depended on the conformational state. We show that a conserved structural motif around T803 with a high similarity to the trypsin catalytic triad is essential for proteolysis. We show that LonP1 is not regulated by redox potential, despite the presence of two conserved cysteines at disulfide-bonding distance in its unfoldase core. Our data indicate how sequential ATP hydrolysis controls substrate protein translocation in a 6-fold binding change mechanism. Substrate protein translocation, rather than ATP hydrolysis, is a rate-limiting step, suggesting that LonP1 is a Brownian ratchet with ATP hydrolysis preventing translocation reversal. 3-fold rocking motions of the flexible N-domain assembly may assist thermal unfolding of the substrate protein. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13102.map.gz emd_13102.map.gz | 229.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13102-v30.xml emd-13102-v30.xml emd-13102.xml emd-13102.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13102.png emd_13102.png | 195.3 KB | ||
| Filedesc metadata |  emd-13102.cif.gz emd-13102.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13102 http://ftp.pdbj.org/pub/emdb/structures/EMD-13102 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13102 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13102 | HTTPS FTP |
-Related structure data
| Related structure data |  7oxoMC  7nfyC  7ng4C  7ng5C  7ngcC  7ngfC  7nglC  7ngpC  7ngqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13102.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13102.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LonP1, R-state, ADP bound, incubated in AMPPCP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.058 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
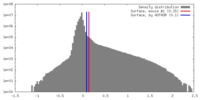
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human mitochondrial Lon protease homolog
| Entire | Name: human mitochondrial Lon protease homolog |
|---|---|
| Components |
|
-Supramolecule #1: human mitochondrial Lon protease homolog
| Supramolecule | Name: human mitochondrial Lon protease homolog / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 639 KDa |
-Macromolecule #1: Lon protease homolog, mitochondrial
| Macromolecule | Name: Lon protease homolog, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: endopeptidase La |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 106.635375 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAASTGYVRL WGAARCWVLR RPMLAAAGGR VPTAAGAWLL RGQRTCDASP PWALWGRGPA IGGQWRGFWE ASSRGGGAFS GGEDASEGG AEEGAGGAGG SAGAGEGPVI TALTPMTIPD VFPHLPLIAI TRNPVFPRFI KIIEVKNKKL VELLRRKVRL A QPYVGVFL ...String: MAASTGYVRL WGAARCWVLR RPMLAAAGGR VPTAAGAWLL RGQRTCDASP PWALWGRGPA IGGQWRGFWE ASSRGGGAFS GGEDASEGG AEEGAGGAGG SAGAGEGPVI TALTPMTIPD VFPHLPLIAI TRNPVFPRFI KIIEVKNKKL VELLRRKVRL A QPYVGVFL KRDDSNESDV VESLDEIYHT GTFAQIHEMQ DLGDKLRMIV MGHRRVHISR QLEVEPEEPE AENKHKPRRK SK RGKKEAE DELSARHPAE LAMEPTPELP AEVLMVEVEN VVHEDFQVTE EVKALTAEIV KTIRDIIALN PLYRESVLQM MQA GQRVVD NPIYLSDMGA ALTGAESHEL QDVLEETNIP KRLYKALSLL KKEFELSKLQ QRLGREVEEK IKQTHRKYLL QEQL KIIKK ELGLEKDDKD AIEEKFRERL KELVVPKHVM DVVDEELSKL GLLDNHSSEF NVTRNYLDWL TSIPWGKYSN ENLDL ARAQ AVLEEDHYGM EDVKKRILEF IAVSQLRGST QGKILCFYGP PGVGKTSIAR SIARALNREY FRFSVGGMTD VAEIKG HRR TYVGAMPGKI IQCLKKTKTE NPLILIDEVD KIGRGYQGDP SSALLELLDP EQNANFLDHY LDVPVDLSKV LFICTAN VT DTIPEPLRDR MEMINVSGYV AQEKLAIAER YLVPQARALC GLDESKAKLS SDVLTLLIKQ YCRESGVRNL QKQVEKVL R KSAYKIVSGE AESVEVTPEN LQDFVGKPVF TVERMYDVTP PGVVMGLAWT AMGGSTLFVE TSLRRPQDKD AKGDKDGSL EVTGQLGEVM KESARIAYTF ARAFLMQHAP ANDYLVTSHI HLHVPEGATP KDGPSAGCTI VTALLSLAMG RPVRQNLAMT GEVSLTGKI LPVGGIKEKT IAAKRAGVTC IVLPAENKKD FYDLAAFITE GLEVHFVEHY REIFDIAFPD EQAEALAVER UniProtKB: Lon protease homolog, mitochondrial |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 80.0 e/Å2 Details: Dat collected in movie mode, 79850 particles used for map reconstruction |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)