[English] 日本語
 Yorodumi
Yorodumi- EMDB-12956: Lassa virus L protein in an elongation conformation [ELONGATION] -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12956 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lassa virus L protein in an elongation conformation [ELONGATION] | |||||||||
 Map data Map data | Lassa virus L protein in an elongation conformation [ELONGATION] | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Lassa virus RNA-dependent RNA polymerase viral RNA / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative stranded viral RNA replication / cap snatching / virion component / host cell / Hydrolases; Acting on ester bonds / host cell cytoplasm / RNA-directed RNA polymerase / nucleotide binding / hydrolase activity / RNA-directed RNA polymerase activity / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Lassa mammarenavirus Lassa mammarenavirus | |||||||||
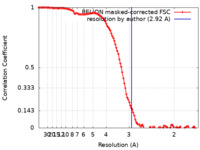
| Method | single particle reconstruction / cryo EM / Resolution: 2.92 Å | |||||||||
 Authors Authors | Kouba T / Vogel D | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Conformational changes in Lassa virus L protein associated with promoter binding and RNA synthesis activity. Authors: Tomas Kouba / Dominik Vogel / Sigurdur R Thorkelsson / Emmanuelle R J Quemin / Harry M Williams / Morlin Milewski / Carola Busch / Stephan Günther / Kay Grünewald / Maria Rosenthal / Stephen Cusack /   Abstract: Lassa virus is endemic in West Africa and can cause severe hemorrhagic fever. The viral L protein transcribes and replicates the RNA genome via its RNA-dependent RNA polymerase activity. Here, we ...Lassa virus is endemic in West Africa and can cause severe hemorrhagic fever. The viral L protein transcribes and replicates the RNA genome via its RNA-dependent RNA polymerase activity. Here, we present nine cryo-EM structures of the L protein in the apo-, promoter-bound pre-initiation and active RNA synthesis states. We characterize distinct binding pockets for the conserved 3' and 5' promoter RNAs and show how full-promoter binding induces a distinct pre-initiation conformation. In the apo- and early elongation states, the endonuclease is inhibited by two distinct L protein peptides, whereas in the pre-initiation state it is uninhibited. In the early elongation state, a template-product duplex is bound in the active site cavity together with an incoming non-hydrolysable nucleotide and the full C-terminal region of the L protein, including the putative cap-binding domain, is well-ordered. These data advance our mechanistic understanding of how this flexible and multifunctional molecular machine is activated. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12956.map.gz emd_12956.map.gz | 61.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12956-v30.xml emd-12956-v30.xml emd-12956.xml emd-12956.xml | 28.2 KB 28.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12956_fsc.xml emd_12956_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_12956.png emd_12956.png | 100.5 KB | ||
| Filedesc metadata |  emd-12956.cif.gz emd-12956.cif.gz | 8.5 KB | ||
| Others |  emd_12956_additional_1.map.gz emd_12956_additional_1.map.gz emd_12956_half_map_1.map.gz emd_12956_half_map_1.map.gz emd_12956_half_map_2.map.gz emd_12956_half_map_2.map.gz | 117.3 MB 98.4 MB 98.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12956 http://ftp.pdbj.org/pub/emdb/structures/EMD-12956 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12956 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12956 | HTTPS FTP |
-Related structure data
| Related structure data |  7ojnMC  7ochC  7oe3C  7oe7C  7oeaC  7oebC  7ojjC  7ojkC  7ojlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12956.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12956.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Lassa virus L protein in an elongation conformation [ELONGATION] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_12956_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
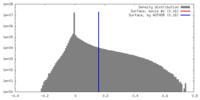
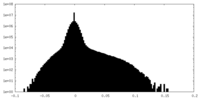
| Density Histograms |
-Half map: #1
| File | emd_12956_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_12956_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : RNA-directed RNA polymerase L Lassa mammarenavirus
+Supramolecule #1: RNA-directed RNA polymerase L Lassa mammarenavirus
+Supramolecule #2: RNA-directed RNA polymerase L
+Supramolecule #3: 5' RNA, 3' RNA
+Supramolecule #4: product RNA
+Macromolecule #1: RNA-directed RNA polymerase L
+Macromolecule #2: 5' RNA
+Macromolecule #3: 3' RNA
+Macromolecule #4: product RNA
+Macromolecule #5: 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phos...
+Macromolecule #6: MANGANESE (II) ION
+Macromolecule #7: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 49.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)







































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)