+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12891 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tetanus neurotoxin HC domain in complex with TT104-Fab1 | |||||||||
 Map data Map data | Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | tetanus / tetanus neurotoxin / humAbs / monoclonal antibody / tetanus prophylaxis / spastic paralysis / tetanus immunoglobulin / TIG / TOXIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationtransmembrane protein transporter activity / metalloendopeptidase activity / toxin activity / proteolysis / extracellular region / zinc ion binding Similarity search - Function | |||||||||
| Biological species |  Clostridium tetani (bacteria) / Clostridium tetani (bacteria) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.0 Å | |||||||||
 Authors Authors | Grinzato A / Kandiah E | |||||||||
 Citation Citation |  Journal: J Clin Invest / Year: 2021 Journal: J Clin Invest / Year: 2021Title: Exceptionally potent human monoclonal antibodies are effective for prophylaxis and treatment of tetanus in mice. Authors: Marco Pirazzini / Alessandro Grinzato / Davide Corti / Sonia Barbieri / Oneda Leka / Francesca Vallese / Marika Tonellato / Chiara Silacci-Fregni / Luca Piccoli / Eaazhisai Kandiah / ...Authors: Marco Pirazzini / Alessandro Grinzato / Davide Corti / Sonia Barbieri / Oneda Leka / Francesca Vallese / Marika Tonellato / Chiara Silacci-Fregni / Luca Piccoli / Eaazhisai Kandiah / Giampietro Schiavo / Giuseppe Zanotti / Antonio Lanzavecchia / Cesare Montecucco /     Abstract: We used human monoclonal antibodies (humAbs) to study the mechanism of neuron intoxication by tetanus neurotoxin and to evaluate these antibodies as a safe preventive and therapeutic substitute for ...We used human monoclonal antibodies (humAbs) to study the mechanism of neuron intoxication by tetanus neurotoxin and to evaluate these antibodies as a safe preventive and therapeutic substitute for hyperimmune sera to treat tetanus in mice. By screening memory B cells from immune donors, we selected 2 tetanus neurotoxin-specific mAbs with exceptionally high neutralizing activities and extensively characterized them both structurally and functionally. We found that these antibodies interfered with the binding and translocation of the neurotoxin into neurons by interacting with 2 epitopes, whose identification pinpoints crucial events in the cellular pathogenesis of tetanus. Our observations explain the neutralization ability of these antibodies, which we found to be exceptionally potent in preventing experimental tetanus when injected into mice long before the toxin. Moreover, their Fab derivatives neutralized tetanus neurotoxin in post-exposure experiments, suggesting their potential for therapeutic use via intrathecal injection. As such, we believe these humAbs, as well as their Fab derivatives, meet the requirements to be considered for prophylactic and therapeutic use in human tetanus and are ready for clinical trials. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12891.map.gz emd_12891.map.gz | 97.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12891-v30.xml emd-12891-v30.xml emd-12891.xml emd-12891.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12891.png emd_12891.png | 81.8 KB | ||
| Filedesc metadata |  emd-12891.cif.gz emd-12891.cif.gz | 5.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12891 http://ftp.pdbj.org/pub/emdb/structures/EMD-12891 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12891 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12891 | HTTPS FTP |
-Related structure data
| Related structure data |  7oh1MC  7oh0C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12891.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12891.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.827 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
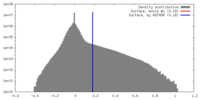
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1
| Entire | Name: Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1 |
|---|---|
| Components |
|
-Supramolecule #1: Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1
| Supramolecule | Name: Tetanus neurotoxin LC-HN domain in complex with TT110-Fab1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Tetanus toxin
| Supramolecule | Name: Tetanus toxin / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Clostridium tetani (bacteria) Clostridium tetani (bacteria) |
-Supramolecule #3: FAB TT110
| Supramolecule | Name: FAB TT110 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Tetanus toxin
| Macromolecule | Name: Tetanus toxin / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Clostridium tetani (bacteria) Clostridium tetani (bacteria) |
| Molecular weight | Theoretical: 99.92432 KDa |
| Sequence | String: MPITINNFRY SDPVNNDTII MMEPPYCKGL DIYYKAFKIT DRIWIVPERY EFGTKPEDFN PPSSLIEGAS EYYDPNYLRT DSDKDRFLQ TMVKLFNRIK NNVAGEALLD KIINAIPYLG NSYSLLDKFD TNSNSVSFNL LEQDPSGATT KSAMLTNLII F GPGPVLNK ...String: MPITINNFRY SDPVNNDTII MMEPPYCKGL DIYYKAFKIT DRIWIVPERY EFGTKPEDFN PPSSLIEGAS EYYDPNYLRT DSDKDRFLQ TMVKLFNRIK NNVAGEALLD KIINAIPYLG NSYSLLDKFD TNSNSVSFNL LEQDPSGATT KSAMLTNLII F GPGPVLNK NEVRGIVLRV DNKNYFPCRD GFGSIMQMAF CPEYVPTFDN VIENITSLTI GKSKYFQDPA LLLMHELIHV LH GLYGMQV SSHEIIPSKQ EIYMQHTYPI SAEELFTFGG QDANLISIDI KNDLYEKTLN DYKAIANKLS QVTSCNDPNI DID SYKQIY QQKYQFDKDS NGQYIVNEDK FQILYNSIMY GFTEIELGKK FNIKTRLSYF SMNHDPVKIP NLLDDTIYND TEGF NIESK DLKSEYKGQN MRVNTNAFRN VDGSGLVSKL IGLCKKIIPP TNIRENLYNR TASLTDLGGE LCIKIKNEDL TFIAE KNSF SEEPFQDEIV SYNTKNKPLN FNYSLDKIIV DYNLQSKITL PNDRTTPVTK GIPYAPEYKS NAASTIEIHN IDDNTI YQY LYAQKSPTTL QRITMTNSVD DALINSTKIY SYFPSVISKV NQGAQGILFL QWVRDIIDDF TNESSQKTTI DKISDVS TI VPYIGPALNI VKQGYEGNFI GALETTGVVL LLEYIPEITL PVIAALSIAE SSTQKEKIIK TIDNFLEKRY EKWIEVYK L VKAKWLGTVN TQFQKRSYQM YRSLEYQVDA IKKIIDYEYK IYSGPDKEQI ADEINNLKNK LEEKANKAMI NINIFMRES SRSFLVNQMI NEAKKQLLEF DTQSKNILMQ YIKANSKFIG ITELKKLESK INKVFSTPIP FSYSKNLDCW UniProtKB: Tetanus toxin |
-Macromolecule #2: FAB TT110
| Macromolecule | Name: FAB TT110 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.298252 KDa |
| Sequence | String: QVQLMQSGAE VQKPGASVKV SCQASGFTLN NYYIHWLRQA PGQGFEWMGI FNPSSGTRTY AQKFQGRISM TADASTTTLY MELSGLTSE DAGVYFCARI GGSTYGRLMT YYFDHWGQGT VVAVSSASTK GPSVFPLAPS SKSTSGGTAA LGCLVKDYFP E PVTVSWNS ...String: QVQLMQSGAE VQKPGASVKV SCQASGFTLN NYYIHWLRQA PGQGFEWMGI FNPSSGTRTY AQKFQGRISM TADASTTTLY MELSGLTSE DAGVYFCARI GGSTYGRLMT YYFDHWGQGT VVAVSSASTK GPSVFPLAPS SKSTSGGTAA LGCLVKDYFP E PVTVSWNS GALTSGVHTF PAVLQSSGLY SLSSVVTVPS SSLGTQTYIC NVNHKPSNTK VDKRVEPKSC |
-Macromolecule #3: FAB TT110
| Macromolecule | Name: FAB TT110 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.023553 KDa |
| Sequence | String: VLTQGPVTLS VSPGGRGTLS CRASRSISTT LAWYQQKPGQ APRLLIYGAS TRATGIPARF TGSGSGTEFT LTISSLQSED FAVYYCQQY NDWPVTFGQG TQVEVKRTVA APSVFIFPPS DEQLKSGTAS VVCLLNNFYP REAKVQWKVD NALQSGNSQE S VTEQDSKD ...String: VLTQGPVTLS VSPGGRGTLS CRASRSISTT LAWYQQKPGQ APRLLIYGAS TRATGIPARF TGSGSGTEFT LTISSLQSED FAVYYCQQY NDWPVTFGQG TQVEVKRTVA APSVFIFPPS DEQLKSGTAS VVCLLNNFYP REAKVQWKVD NALQSGNSQE S VTEQDSKD STYSLSSTLT LSKADYEKHK VYACEVTHQG LSSPVTKSFN RGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 8.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 98170 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)