[English] 日本語
 Yorodumi
Yorodumi- EMDB-11319: The pointed end complex of dynactin with the p150 projection docked -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11319 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The pointed end complex of dynactin with the p150 projection docked | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dynactin / Complex / Scaffold / Cytoskeleton / STRUCTURAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationretrograde axonal transport of mitochondrion / Gap junction degradation / Formation of annular gap junctions / Regulation of actin dynamics for phagocytic cup formation / EPHB-mediated forward signaling / Adherens junctions interactions / VEGFA-VEGFR2 Pathway / Cell-extracellular matrix interactions / RHO GTPases Activate WASPs and WAVEs / MAP2K and MAPK activation ...retrograde axonal transport of mitochondrion / Gap junction degradation / Formation of annular gap junctions / Regulation of actin dynamics for phagocytic cup formation / EPHB-mediated forward signaling / Adherens junctions interactions / VEGFA-VEGFR2 Pathway / Cell-extracellular matrix interactions / RHO GTPases Activate WASPs and WAVEs / MAP2K and MAPK activation / UCH proteinases / Clathrin-mediated endocytosis / RHOF GTPase cycle / dynactin complex / Regulation of PLK1 Activity at G2/M Transition / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Anchoring of the basal body to the plasma membrane / AURKA Activation by TPX2 / Recruitment of mitotic centrosome proteins and complexes / structural constituent of postsynaptic actin cytoskeleton / dense body / dynein complex / Neutrophil degranulation / RHO GTPases activate IQGAPs / RHO GTPases Activate Formins / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / MHC class II antigen presentation / Recruitment of NuMA to mitotic centrosomes / COPI-mediated anterograde transport / NuA4 histone acetyltransferase complex / dynein complex binding / microtubule-based process / stress fiber / axon cytoplasm / sarcomere / axonogenesis / mitotic spindle organization / cell motility / actin filament / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / kinetochore / actin cytoskeleton / cell cortex / microtubule / cytoskeleton / hydrolase activity / axon / focal adhesion / centrosome / synapse / protein kinase binding / protein-containing complex / ATP binding / membrane / nucleus / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
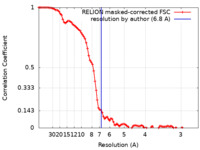
| Method | single particle reconstruction / cryo EM / Resolution: 6.8 Å | |||||||||
 Authors Authors | Lau CK / Lacey SE | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2021 Journal: EMBO J / Year: 2021Title: Cryo-EM reveals the complex architecture of dynactin's shoulder region and pointed end. Authors: Clinton K Lau / Francis J O'Reilly / Balaji Santhanam / Samuel E Lacey / Juri Rappsilber / Andrew P Carter /   Abstract: Dynactin is a 1.1 MDa complex that activates the molecular motor dynein for ultra-processive transport along microtubules. In order to do this, it forms a tripartite complex with dynein and a coiled- ...Dynactin is a 1.1 MDa complex that activates the molecular motor dynein for ultra-processive transport along microtubules. In order to do this, it forms a tripartite complex with dynein and a coiled-coil adaptor. Dynactin consists of an actin-related filament whose length is defined by its flexible shoulder domain. Despite previous cryo-EM structures, the molecular architecture of the shoulder and pointed end of the filament is still poorly understood due to the lack of high-resolution information in these regions. Here we combine multiple cryo-EM datasets and define precise masking strategies for particle signal subtraction and 3D classification. This overcomes domain flexibility and results in high-resolution maps into which we can build the shoulder and pointed end. The unique architecture of the shoulder securely houses the p150 subunit and positions the four identical p50 subunits in different conformations to bind dynactin's filament. The pointed end map allows us to build the first structure of p62 and reveals the molecular basis for cargo adaptor binding to different sites at the pointed end. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11319.map.gz emd_11319.map.gz | 60.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11319-v30.xml emd-11319-v30.xml emd-11319.xml emd-11319.xml | 27 KB 27 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11319_fsc.xml emd_11319_fsc.xml | 9.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_11319.png emd_11319.png | 40.1 KB | ||
| Masks |  emd_11319_msk_1.map emd_11319_msk_1.map | 67 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11319.cif.gz emd-11319.cif.gz | 7.2 KB | ||
| Others |  emd_11319_half_map_1.map.gz emd_11319_half_map_1.map.gz emd_11319_half_map_2.map.gz emd_11319_half_map_2.map.gz | 52.1 MB 52.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11319 http://ftp.pdbj.org/pub/emdb/structures/EMD-11319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11319 | HTTPS FTP |
-Validation report
| Summary document |  emd_11319_validation.pdf.gz emd_11319_validation.pdf.gz | 803.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11319_full_validation.pdf.gz emd_11319_full_validation.pdf.gz | 803.1 KB | Display | |
| Data in XML |  emd_11319_validation.xml.gz emd_11319_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_11319_validation.cif.gz emd_11319_validation.cif.gz | 21.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11319 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11319 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11319 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11319 | HTTPS FTP |
-Related structure data
| Related structure data |  6znoMC  6znlC  6znmC  6znnC  6zo4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11319.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11319.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11319_msk_1.map emd_11319_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_11319_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_11319_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Dynactin
+Supramolecule #1: Dynactin
+Macromolecule #1: ARP1 actin related protein 1 homolog A
+Macromolecule #2: Actin, cytoplasmic 1
+Macromolecule #3: Arp11
+Macromolecule #4: Dynactin subunit 2
+Macromolecule #5: Dynactin 6
+Macromolecule #6: Dynactin subunit 5
+Macromolecule #7: Dynactin subunit 4
+Macromolecule #8: Dynactin subunit 1
+Macromolecule #9: Dynactin subunit 1
+Macromolecule #10: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #11: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #12: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: FEI FALCON II (4k x 4k) / #0 - Average electron dose: 52.0 e/Å2 / #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K3 (6k x 4k) / #1 - Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)