+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11243 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Membrane domain of closed complex I during turnover | |||||||||
 Map data Map data | Oversampled, local resolution-filtered map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationNADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / : / mitochondrial electron transport, NADH to ubiquinone / mitochondrial respiratory chain complex I assembly / electron transport coupled proton transport / acyl binding / acyl carrier activity / NADH dehydrogenase (ubiquinone) activity / : ...NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / : / mitochondrial electron transport, NADH to ubiquinone / mitochondrial respiratory chain complex I assembly / electron transport coupled proton transport / acyl binding / acyl carrier activity / NADH dehydrogenase (ubiquinone) activity / : / ATP synthesis coupled electron transport / reactive oxygen species metabolic process / electron transport chain / mitochondrial intermembrane space / membrane => GO:0016020 / mitochondrial inner membrane / mitochondrial matrix Similarity search - Function | |||||||||
| Biological species |   | |||||||||
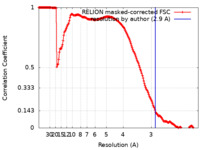
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Kampjut D / Sazanov LA | |||||||||
| Funding support | European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2020 Journal: Science / Year: 2020Title: The coupling mechanism of mammalian respiratory complex I. Authors: Domen Kampjut / Leonid A Sazanov /  Abstract: Mitochondrial complex I couples NADH:ubiquinone oxidoreduction to proton pumping by an unknown mechanism. Here, we present cryo-electron microscopy structures of ovine complex I in five different ...Mitochondrial complex I couples NADH:ubiquinone oxidoreduction to proton pumping by an unknown mechanism. Here, we present cryo-electron microscopy structures of ovine complex I in five different conditions, including turnover, at resolutions up to 2.3 to 2.5 angstroms. Resolved water molecules allowed us to experimentally define the proton translocation pathways. Quinone binds at three positions along the quinone cavity, as does the inhibitor rotenone that also binds within subunit ND4. Dramatic conformational changes around the quinone cavity couple the redox reaction to proton translocation during open-to-closed state transitions of the enzyme. In the induced deactive state, the open conformation is arrested by the ND6 subunit. We propose a detailed molecular coupling mechanism of complex I, which is an unexpected combination of conformational changes and electrostatic interactions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11243.map.gz emd_11243.map.gz | 149.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11243-v30.xml emd-11243-v30.xml emd-11243.xml emd-11243.xml | 40.4 KB 40.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11243_fsc.xml emd_11243_fsc.xml | 17.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_11243.png emd_11243.png | 192.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11243 http://ftp.pdbj.org/pub/emdb/structures/EMD-11243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11243 | HTTPS FTP |
-Validation report
| Summary document |  emd_11243_validation.pdf.gz emd_11243_validation.pdf.gz | 284.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11243_full_validation.pdf.gz emd_11243_full_validation.pdf.gz | 283.7 KB | Display | |
| Data in XML |  emd_11243_validation.xml.gz emd_11243_validation.xml.gz | 12.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11243 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11243 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11243 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11243 | HTTPS FTP |
-Related structure data
| Related structure data |  6zkbMC  6zk9C  6zkaC  6zkcC  6zkdC  6zkeC  6zkfC  6zkgC  6zkhC  6zkiC  6zkjC  6zkkC  6zklC  6zkmC  6zknC  6zkoC  6zkpC  6zkqC  6zkrC  6zksC  6zktC  6zkuC  6zkvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11243.map.gz / Format: CCP4 / Size: 163.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11243.map.gz / Format: CCP4 / Size: 163.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Oversampled, local resolution-filtered map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
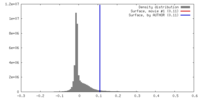
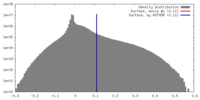
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Membrane domain of closed complex I during turnover
+Supramolecule #1: Membrane domain of closed complex I during turnover
+Macromolecule #1: Mitochondrial complex I, 49 kDa subunit
+Macromolecule #2: NADH-ubiquinone oxidoreductase chain 5
+Macromolecule #3: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #4: NADH-ubiquinone oxidoreductase chain 2
+Macromolecule #5: Mitochondrial complex I, B14.7 subunit
+Macromolecule #6: NADH:ubiquinone oxidoreductase subunit B5
+Macromolecule #7: Acyl carrier protein
+Macromolecule #8: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 8
+Macromolecule #9: Mitochondrial complex I, PDSW subunit
+Macromolecule #10: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 10, mi...
+Macromolecule #11: NADH:ubiquinone oxidoreductase subunit S5
+Macromolecule #12: NADH:ubiquinone oxidoreductase subunit A3
+Macromolecule #13: NADH:ubiquinone oxidoreductase subunit B3
+Macromolecule #14: NADH dehydrogenase [ubiquinone] 1 subunit C2
+Macromolecule #15: NADH:ubiquinone oxidoreductase subunit B4
+Macromolecule #16: Mitochondrial complex I, B16.6 subunit
+Macromolecule #17: Mitochondrial complex I, B17 subunit
+Macromolecule #18: NADH:ubiquinone oxidoreductase subunit B7
+Macromolecule #19: NADH:ubiquinone oxidoreductase subunit B9
+Macromolecule #20: NADH:ubiquinone oxidoreductase subunit B2
+Macromolecule #21: NADH dehydrogenase [ubiquinone] 1 beta subcomplex subunit 8, mito...
+Macromolecule #22: Mitochondrial complex I, ESSS subunit
+Macromolecule #23: Mitochondrial complex I, KFYI subunit
+Macromolecule #24: Mitochondrial complex I, MNLL subunit
+Macromolecule #25: Mitochondrial complex I, MWFE subunit
+Macromolecule #26: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #27: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #28: NADH-ubiquinone oxidoreductase chain 6
+Macromolecule #29: NADH-ubiquinone oxidoreductase chain 4L
+Macromolecule #30: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOETHANOLAMINE
+Macromolecule #31: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #32: CARDIOLIPIN
+Macromolecule #33: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #34: ADENOSINE MONOPHOSPHATE
+Macromolecule #35: MYRISTIC ACID
+Macromolecule #36: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)