+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11209 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GLIC pentameric ligand-gated ion channel, pH 3 | |||||||||
 Map data Map data | GLIC, pentameric ligand-gated ion channel, pH 3, sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GLIC / pentameric ligand-gated ion channel / cryoem / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsodium channel activity / potassium channel activity / extracellular ligand-gated monoatomic ion channel activity / transmembrane signaling receptor activity / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria) Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria) | |||||||||
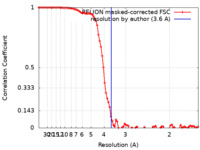
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Rovsnik U / Zhuang Y | |||||||||
| Funding support |  Sweden, 2 items Sweden, 2 items
| |||||||||
 Citation Citation |  Journal: Life Sci Alliance / Year: 2021 Journal: Life Sci Alliance / Year: 2021Title: Dynamic closed states of a ligand-gated ion channel captured by cryo-EM and simulations. Authors: Urška Rovšnik / Yuxuan Zhuang / Björn O Forsberg / Marta Carroni / Linnea Yvonnesdotter / Rebecca J Howard / Erik Lindahl /   Abstract: Ligand-gated ion channels are critical mediators of electrochemical signal transduction across evolution. Biophysical and pharmacological characterization of these receptor proteins relies on high- ...Ligand-gated ion channels are critical mediators of electrochemical signal transduction across evolution. Biophysical and pharmacological characterization of these receptor proteins relies on high-quality structures in multiple, subtly distinct functional states. However, structural data in this family remain limited, particularly for resting and intermediate states on the activation pathway. Here, we report cryo-electron microscopy (cryo-EM) structures of the proton-activated ligand-gated ion channel (GLIC) under three pH conditions. Decreased pH was associated with improved resolution and side chain rearrangements at the subunit/domain interface, particularly involving functionally important residues in the β1-β2 and M2-M3 loops. Molecular dynamics simulations substantiated flexibility in the closed-channel extracellular domains relative to the transmembrane ones and supported electrostatic remodeling around E35 and E243 in proton-induced gating. Exploration of secondary cryo-EM classes further indicated a low-pH population with an expanded pore. These results allow us to define distinct protonation and activation steps in pH-stimulated conformational cycling in GLIC, including interfacial rearrangements largely conserved in the pentameric channel family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11209.map.gz emd_11209.map.gz | 7.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11209-v30.xml emd-11209-v30.xml emd-11209.xml emd-11209.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11209_fsc.xml emd_11209_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_11209.png emd_11209.png | 150.3 KB | ||
| Masks |  emd_11209_msk_1.map emd_11209_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11209.cif.gz emd-11209.cif.gz | 5.8 KB | ||
| Others |  emd_11209_additional_1.map.gz emd_11209_additional_1.map.gz emd_11209_half_map_1.map.gz emd_11209_half_map_1.map.gz emd_11209_half_map_2.map.gz emd_11209_half_map_2.map.gz | 49.6 MB 49.8 MB 49.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11209 http://ftp.pdbj.org/pub/emdb/structures/EMD-11209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11209 | HTTPS FTP |
-Related structure data
| Related structure data |  6zgkMC  6zgdC  6zgjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11209.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11209.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GLIC, pentameric ligand-gated ion channel, pH 3, sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11209_msk_1.map emd_11209_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: GLIC, pentameric ligand-gated ion channel, pH 3, unsharpened map
| File | emd_11209_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GLIC, pentameric ligand-gated ion channel, pH 3, unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: GLIC, pentameric ligand-gated ion channel, pH 3, half map 1
| File | emd_11209_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GLIC, pentameric ligand-gated ion channel, pH 3, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: GLIC, pentameric ligand-gated ion channel, pH 3, half map 2
| File | emd_11209_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | GLIC, pentameric ligand-gated ion channel, pH 3, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Pentameric ion channel
| Entire | Name: Pentameric ion channel |
|---|---|
| Components |
|
-Supramolecule #1: Pentameric ion channel
| Supramolecule | Name: Pentameric ion channel / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria) Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria) |
| Molecular weight | Theoretical: 180 KDa |
-Macromolecule #1: Proton-gated ion channel
| Macromolecule | Name: Proton-gated ion channel / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria) Gloeobacter violaceus (strain ATCC 29082 / PCC 7421) (bacteria)Strain: ATCC 29082 / PCC 7421 |
| Molecular weight | Theoretical: 36.29175 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AQDMVSPPPP IADEPLTVNT GIYLIECYSL DDKAETFKVN AFLSLSWKDR RLAFDPVRSG VRVKTYEPEA IWIPEIRFVN VENARDADV VDISVSPDGT VQYLERFSAR VLSPLDFRRY PFDSQTLHIY LIVRSVDTRN IVLAVDLEKV GKNDDVFLTG W DIESFTAV ...String: AQDMVSPPPP IADEPLTVNT GIYLIECYSL DDKAETFKVN AFLSLSWKDR RLAFDPVRSG VRVKTYEPEA IWIPEIRFVN VENARDADV VDISVSPDGT VQYLERFSAR VLSPLDFRRY PFDSQTLHIY LIVRSVDTRN IVLAVDLEKV GKNDDVFLTG W DIESFTAV VKPANFALED RLESKLDYQL RISRQYFSYI PNIILPMLFI LFISWTAFWS TSYEANVTLV VSTLIAHIAF NI LVETNLP KTPYMTYTGA IIFMIYLFYF VAVIEVTVQH YLKVESQPAR AASITRASRI AFPVVFLLAN IILAFLFFGF UniProtKB: Proton-gated ion channel |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 3 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 1.5 s force -1. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80.0 K |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-40 / Number grids imaged: 1 / Number real images: 6000 / Average exposure time: 6.0 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 0.0028 µm / Calibrated defocus min: 0.0009000000000000001 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.8000000000000003 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)