[English] 日本語
 Yorodumi
Yorodumi- EMDB-10360: Bat Influenza A polymerase elongation complex with incoming UTP a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10360 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bat Influenza A polymerase elongation complex with incoming UTP analogue (complete polymerase) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Influenza / polymerase / viral transcription / RNA / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / host cell cytoplasm / symbiont-mediated suppression of host gene expression / viral translational frameshifting / RNA-directed RNA polymerase ...cap snatching / viral transcription / symbiont-mediated suppression of host mRNA transcription via inhibition of RNA polymerase II activity / host cell mitochondrion / 7-methylguanosine mRNA capping / virion component / host cell cytoplasm / symbiont-mediated suppression of host gene expression / viral translational frameshifting / RNA-directed RNA polymerase / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / GTP binding / host cell nucleus / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Influenza A virus (A/little yellow-shouldered bat/Guatemala/060/2010(H17N10)) Influenza A virus (A/little yellow-shouldered bat/Guatemala/060/2010(H17N10)) | |||||||||
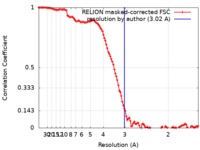
| Method | single particle reconstruction / cryo EM / Resolution: 3.02 Å | |||||||||
 Authors Authors | Wandzik JM / Kouba T | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: A Structure-Based Model for the Complete Transcription Cycle of Influenza Polymerase. Authors: Joanna M Wandzik / Tomas Kouba / Manikandan Karuppasamy / Alexander Pflug / Petra Drncova / Jan Provaznik / Nayara Azevedo / Stephen Cusack /   Abstract: Influenza polymerase uses unique mechanisms to synthesize capped and polyadenylated mRNAs from the genomic viral RNA (vRNA) template, which is packaged inside ribonucleoprotein particles (vRNPs). ...Influenza polymerase uses unique mechanisms to synthesize capped and polyadenylated mRNAs from the genomic viral RNA (vRNA) template, which is packaged inside ribonucleoprotein particles (vRNPs). Here, we visualize by cryoelectron microscopy the conformational dynamics of the polymerase during the complete transcription cycle from pre-initiation to termination, focusing on the template trajectory. After exiting the active site cavity, the template 3' extremity rebinds into a specific site on the polymerase surface. Here, it remains sequestered during all subsequent transcription steps, forcing the template to loop out as it further translocates. At termination, the strained connection between the bound template 5' end and the active site results in polyadenylation by stuttering at uridine 17. Upon product dissociation, further conformational changes release the trapped template, allowing recycling back into the pre-initiation state. Influenza polymerase thus performs transcription while tightly binding to and protecting both template ends, allowing efficient production of multiple mRNAs from a single vRNP. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10360.map.gz emd_10360.map.gz | 117.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10360-v30.xml emd-10360-v30.xml emd-10360.xml emd-10360.xml | 26.2 KB 26.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10360_fsc.xml emd_10360_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_10360.png emd_10360.png | 176.8 KB | ||
| Filedesc metadata |  emd-10360.cif.gz emd-10360.cif.gz | 8.2 KB | ||
| Others |  emd_10360_half_map_1.map.gz emd_10360_half_map_1.map.gz emd_10360_half_map_2.map.gz emd_10360_half_map_2.map.gz | 98.5 MB 98.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10360 http://ftp.pdbj.org/pub/emdb/structures/EMD-10360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10360 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10360 | HTTPS FTP |
-Related structure data
| Related structure data |  6t0vMC  6szuC  6szvC  6t0nC  6t0rC  6t0sC  6t0uC  6t0wC  6t2cC  6tu5C  6tw1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10360.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10360.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8127 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #1
| File | emd_10360_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_10360_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Bat Influenza A polymerase elongation complex with incoming UTP a...
+Supramolecule #1: Bat Influenza A polymerase elongation complex with incoming UTP a...
+Supramolecule #2: Polymerase
+Supramolecule #3: Nucleic acids
+Macromolecule #1: Polymerase acidic protein
+Macromolecule #2: RNA-directed RNA polymerase catalytic subunit
+Macromolecule #3: Polymerase basic protein 2
+Macromolecule #4: 5' vRNA
+Macromolecule #5: 3' vRNA
+Macromolecule #6: Capped mRNA
+Macromolecule #7: 5-oxidanyl-4-oxidanylidene-1-[(1-pyrrolo[2,3-b]pyridin-1-ylcyclop...
+Macromolecule #8: MAGNESIUM ION
+Macromolecule #9: 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phos...
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)

































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)