+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-8665 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Nucleotide-driven Triple-state Remodeling of the AAA-ATPase Channel in the Activated Human 26S Proteasome | |||||||||
 マップデータ マップデータ | Final SD3 state map with a corrected B-factor of -80 and low-pass filtered to 4.9 Angstrom | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | 26S proteasome / ATP-dependent protease / AAA-ATPase / peptide-unfolding channel / 20S core particle / HYDROLASE | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報positive regulation of inclusion body assembly / Impaired BRCA2 translocation to the nucleus / Impaired BRCA2 binding to SEM1 (DSS1) / thyrotropin-releasing hormone receptor binding / modulation by host of viral transcription / meiosis I / 加水分解酵素; プロテアーゼ; ペプチド結合加水分解酵素; オメガペプチターゼ / proteasome accessory complex / integrator complex / purine ribonucleoside triphosphate binding ...positive regulation of inclusion body assembly / Impaired BRCA2 translocation to the nucleus / Impaired BRCA2 binding to SEM1 (DSS1) / thyrotropin-releasing hormone receptor binding / modulation by host of viral transcription / meiosis I / 加水分解酵素; プロテアーゼ; ペプチド結合加水分解酵素; オメガペプチターゼ / proteasome accessory complex / integrator complex / purine ribonucleoside triphosphate binding / proteasome regulatory particle / cytosolic proteasome complex / positive regulation of proteasomal protein catabolic process / proteasome regulatory particle, lid subcomplex / proteasome-activating activity / proteasome regulatory particle, base subcomplex / metal-dependent deubiquitinase activity / negative regulation of programmed cell death / regulation of endopeptidase activity / protein K63-linked deubiquitination / Homologous DNA Pairing and Strand Exchange / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Regulation of ornithine decarboxylase (ODC) / proteasome core complex / Resolution of D-loop Structures through Holliday Junction Intermediates / Cross-presentation of soluble exogenous antigens (endosomes) / : / K63-linked deubiquitinase activity / Somitogenesis / Impaired BRCA2 binding to RAD51 / myofibril / immune system process / proteasome binding / transcription factor binding / regulation of protein catabolic process / proteasome storage granule / Presynaptic phase of homologous DNA pairing and strand exchange / blastocyst development / general transcription initiation factor binding / NF-kappaB binding / endopeptidase activator activity / polyubiquitin modification-dependent protein binding / proteasome assembly / protein deubiquitination / positive regulation of RNA polymerase II transcription preinitiation complex assembly / proteasome endopeptidase complex / proteasome core complex, beta-subunit complex / proteasome core complex, alpha-subunit complex / threonine-type endopeptidase activity / mRNA export from nucleus / regulation of proteasomal protein catabolic process / enzyme regulator activity / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / inclusion body / ERAD pathway / sarcomere / proteasome complex / ciliary basal body / Regulation of activated PAK-2p34 by proteasome mediated degradation / proteolysis involved in protein catabolic process / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / Ubiquitin-dependent degradation of Cyclin D / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / stem cell differentiation / Vpu mediated degradation of CD4 / Assembly of the pre-replicative complex / Degradation of DVL / negative regulation of inflammatory response to antigenic stimulus / Ubiquitin Mediated Degradation of Phosphorylated Cdc25A / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Dectin-1 mediated noncanonical NF-kB signaling / lipopolysaccharide binding / Hh mutants are degraded by ERAD / Degradation of AXIN / Activation of NF-kappaB in B cells / Degradation of GLI1 by the proteasome / Hedgehog ligand biogenesis / Defective CFTR causes cystic fibrosis / G2/M Checkpoints / Negative regulation of NOTCH4 signaling / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Autodegradation of the E3 ubiquitin ligase COP1 / Vif-mediated degradation of APOBEC3G / double-strand break repair via homologous recombination / P-body / Regulation of RUNX3 expression and activity / Hedgehog 'on' state / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / MAPK6/MAPK4 signaling 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.9 Å | |||||||||
 データ登録者 データ登録者 | Zhu Y / Wang WL | |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2018 ジャーナル: Nat Commun / 年: 2018タイトル: Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome. 著者: Yanan Zhu / Wei Li Wang / Daqi Yu / Qi Ouyang / Ying Lu / Youdong Mao /   要旨: The proteasome is a sophisticated ATP-dependent molecular machine responsible for protein degradation in all known eukaryotic cells. It remains elusive how conformational changes of the AAA-ATPase ...The proteasome is a sophisticated ATP-dependent molecular machine responsible for protein degradation in all known eukaryotic cells. It remains elusive how conformational changes of the AAA-ATPase unfoldase in the regulatory particle (RP) control the gating of the substrate-translocation channel leading to the proteolytic chamber of the core particle (CP). Here we report three alternative states of the ATP-γ-S-bound human proteasome, in which the CP gates are asymmetrically open, visualized by cryo-EM at near-atomic resolutions. At least four nucleotides are bound to the AAA-ATPase ring in these open-gate states. Variation in nucleotide binding gives rise to an axial movement of the pore loops narrowing the substrate-translation channel, which exhibit remarkable structural transitions between the spiral-staircase and saddle-shaped-circle topologies. Gate opening in the CP is thus regulated by nucleotide-driven conformational changes of the AAA-ATPase unfoldase. These findings demonstrate an elegant mechanism of allosteric coordination among sub-machines within the human proteasome holoenzyme. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_8665.map.gz emd_8665.map.gz | 612.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-8665-v30.xml emd-8665-v30.xml emd-8665.xml emd-8665.xml | 57.6 KB 57.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_8665.png emd_8665.png | 62.7 KB | ||
| Filedesc metadata |  emd-8665.cif.gz emd-8665.cif.gz | 13.4 KB | ||
| その他 |  emd_8665_additional_1.map.gz emd_8665_additional_1.map.gz emd_8665_additional_2.map.gz emd_8665_additional_2.map.gz | 612.8 MB 593.6 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8665 http://ftp.pdbj.org/pub/emdb/structures/EMD-8665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8665 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_8665_validation.pdf.gz emd_8665_validation.pdf.gz | 689.1 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_8665_full_validation.pdf.gz emd_8665_full_validation.pdf.gz | 688.6 KB | 表示 | |
| XML形式データ |  emd_8665_validation.xml.gz emd_8665_validation.xml.gz | 6.8 KB | 表示 | |
| CIF形式データ |  emd_8665_validation.cif.gz emd_8665_validation.cif.gz | 8.1 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8665 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8665 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8665 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8665 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  5vfrMC  8662C  8663C  8664C  8666C  8667C  8668C  5vfoC  5vfpC  5vfqC  5vfsC  5vftC  5vfuC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10090 (タイトル: Nucleotide-Driven Triple-State Remodeling of the AAA-ATPase Channel in the Activated Human 26S Proteasome EMPIAR-10090 (タイトル: Nucleotide-Driven Triple-State Remodeling of the AAA-ATPase Channel in the Activated Human 26S ProteasomeData size: 2.8 TB Data #1: Motion-corrected single frame micrographs of ATP-gS-bound human proteasome [micrographs - single frame] Data #2: Single-particle stacks of classified conformations [picked particles - multiframe - processed]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_8665.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_8665.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Final SD3 state map with a corrected B-factor of -80 and low-pass filtered to 4.9 Angstrom | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.75 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
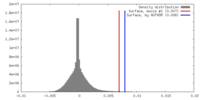
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: Final SD3 state with a local mask of AAA-ATPase and 20S core particle
| ファイル | emd_8665_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Final SD3 state with a local mask of AAA-ATPase and 20S core particle | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-追加マップ: Uncorrected raw map
| ファイル | emd_8665_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Uncorrected raw map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : 26S proteasome bound to ATP-gammaS
+超分子 #1: 26S proteasome bound to ATP-gammaS
+分子 #1: 26S proteasome non-ATPase regulatory subunit 1
+分子 #2: 26S proteasome non-ATPase regulatory subunit 3
+分子 #3: 26S proteasome non-ATPase regulatory subunit 12
+分子 #4: 26S proteasome non-ATPase regulatory subunit 11
+分子 #5: 26S proteasome non-ATPase regulatory subunit 6
+分子 #6: 26S proteasome non-ATPase regulatory subunit 7
+分子 #7: 26S proteasome non-ATPase regulatory subunit 13
+分子 #8: 26S proteasome non-ATPase regulatory subunit 4
+分子 #9: 26S proteasome non-ATPase regulatory subunit 14
+分子 #10: 26S proteasome non-ATPase regulatory subunit 8
+分子 #11: Sem1
+分子 #12: Proteasome subunit alpha type-6
+分子 #13: Proteasome subunit alpha type-2
+分子 #14: Proteasome subunit alpha type-4
+分子 #15: Proteasome subunit alpha type-7
+分子 #16: Proteasome subunit alpha type-5
+分子 #17: Proteasome subunit alpha type-1
+分子 #18: Proteasome subunit alpha type-3
+分子 #19: Proteasome subunit beta type-6
+分子 #20: Proteasome subunit beta type-7
+分子 #21: Proteasome subunit beta type-3
+分子 #22: Proteasome subunit beta type-2
+分子 #23: Proteasome subunit beta type-5
+分子 #24: Proteasome subunit beta type-1
+分子 #25: Proteasome subunit beta type-4
+分子 #26: 26S proteasome regulatory subunit 7
+分子 #27: 26S proteasome regulatory subunit 4
+分子 #28: 26S proteasome regulatory subunit 8
+分子 #29: 26S proteasome regulatory subunit 6B
+分子 #30: 26S proteasome regulatory subunit 10B
+分子 #31: 26S proteasome regulatory subunit 6A
+分子 #32: 26S proteasome non-ATPase regulatory subunit 2
+分子 #33: ZINC ION
+分子 #34: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TECNAI ARCTICA |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 30.0 e/Å2 |
| 電子線 | 加速電圧: 200 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Talos Arctica / 画像提供: FEI Company |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: OTHER |
|---|---|
| 最終 再構成 | 想定した対称性 - 点群: C1 (非対称) / 解像度のタイプ: BY AUTHOR / 解像度: 4.9 Å / 解像度の算出法: FSC 0.143 CUT-OFF / ソフトウェア - 名称: RELION / 使用した粒子像数: 33278 |
| 初期 角度割当 | タイプ: ANGULAR RECONSTITUTION / ソフトウェア - 名称: RELION |
| 最終 角度割当 | タイプ: PROJECTION MATCHING / ソフトウェア - 名称: RELION |
 ムービー
ムービー コントローラー
コントローラー





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)