[English] 日本語
 Yorodumi
Yorodumi- EMDB-8662: Nucleotide-driven Triple-state Remodeling of the AAA-ATPase Chann... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8662 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Nucleotide-driven Triple-state Remodeling of the AAA-ATPase Channel in the Activated Human 26S Proteasome | |||||||||
 Map data Map data | Final map corrected with a B-factor of -80 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | 26S proteasome / ATP-dependent protease / AAA-ATPase / peptide-unfolding channel / 20S core particle / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationpurine ribonucleoside triphosphate binding / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx / Regulation of ornithine decarboxylase (ODC) / Proteasome assembly / proteasome core complex / perinuclear theca / Cross-presentation of soluble exogenous antigens (endosomes) / Somitogenesis / myofibril ...purine ribonucleoside triphosphate binding / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx / Regulation of ornithine decarboxylase (ODC) / Proteasome assembly / proteasome core complex / perinuclear theca / Cross-presentation of soluble exogenous antigens (endosomes) / Somitogenesis / myofibril / proteasomal ubiquitin-independent protein catabolic process / sperm head-tail coupling apparatus / immune system process / proteasome endopeptidase complex / NF-kappaB binding / proteasome core complex, beta-subunit complex / threonine-type endopeptidase activity / proteasome assembly / proteasome core complex, alpha-subunit complex / : / ciliary tip / proteasome complex / Regulation of activated PAK-2p34 by proteasome mediated degradation / sarcomere / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / Ubiquitin-dependent degradation of Cyclin D / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / centriole / sperm end piece / negative regulation of inflammatory response to antigenic stimulus / P-body / Assembly of the pre-replicative complex / Vpu mediated degradation of CD4 / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / lipopolysaccharide binding / Dectin-1 mediated noncanonical NF-kB signaling / Degradation of DVL / Degradation of AXIN / Degradation of CRY and PER proteins / Hh mutants are degraded by ERAD / Activation of NF-kappaB in B cells / G2/M Checkpoints / Degradation of GLI1 by the proteasome / Hedgehog ligand biogenesis / Regulation of RUNX3 expression and activity / Autodegradation of the E3 ubiquitin ligase COP1 / Defective CFTR causes cystic fibrosis / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Negative regulation of NOTCH4 signaling / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Hedgehog 'on' state / Vif-mediated degradation of APOBEC3G / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / MAPK6/MAPK4 signaling / Degradation of CDH1 / Degradation of beta-catenin by the destruction complex / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / CDK-mediated phosphorylation and removal of Cdc6 / ABC-family protein mediated transport / CLEC7A (Dectin-1) signaling / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation / response to virus / Regulation of expression of SLITs and ROBOs / Regulation of PTEN stability and activity / nuclear matrix / Interleukin-1 signaling / Orc1 removal from chromatin / Regulation of RAS by GAPs / Regulation of RUNX2 expression and activity / The role of GTSE1 in G2/M progression after G2 checkpoint / Separation of Sister Chromatids / UCH proteinases / KEAP1-NFE2L2 pathway / Downstream TCR signaling / peptidase activity / Antigen processing: Ubiquitination & Proteasome degradation / sperm principal piece / RUNX1 regulates transcription of genes involved in differentiation of HSCs / ER-Phagosome pathway / Neddylation / regulation of inflammatory response / response to oxidative stress / sperm midpiece / secretory granule lumen / endopeptidase activity / ficolin-1-rich granule lumen / proteasome-mediated ubiquitin-dependent protein catabolic process / positive regulation of canonical NF-kappaB signal transduction / Ub-specific processing proteases / cilium / nuclear body Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zhu Y / Wang WL | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2018 Journal: Nat Commun / Year: 2018Title: Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome. Authors: Yanan Zhu / Wei Li Wang / Daqi Yu / Qi Ouyang / Ying Lu / Youdong Mao /   Abstract: The proteasome is a sophisticated ATP-dependent molecular machine responsible for protein degradation in all known eukaryotic cells. It remains elusive how conformational changes of the AAA-ATPase ...The proteasome is a sophisticated ATP-dependent molecular machine responsible for protein degradation in all known eukaryotic cells. It remains elusive how conformational changes of the AAA-ATPase unfoldase in the regulatory particle (RP) control the gating of the substrate-translocation channel leading to the proteolytic chamber of the core particle (CP). Here we report three alternative states of the ATP-γ-S-bound human proteasome, in which the CP gates are asymmetrically open, visualized by cryo-EM at near-atomic resolutions. At least four nucleotides are bound to the AAA-ATPase ring in these open-gate states. Variation in nucleotide binding gives rise to an axial movement of the pore loops narrowing the substrate-translation channel, which exhibit remarkable structural transitions between the spiral-staircase and saddle-shaped-circle topologies. Gate opening in the CP is thus regulated by nucleotide-driven conformational changes of the AAA-ATPase unfoldase. These findings demonstrate an elegant mechanism of allosteric coordination among sub-machines within the human proteasome holoenzyme. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8662.map.gz emd_8662.map.gz | 613.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8662-v30.xml emd-8662-v30.xml emd-8662.xml emd-8662.xml | 31.4 KB 31.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8662.png emd_8662.png | 82.4 KB | ||
| Filedesc metadata |  emd-8662.cif.gz emd-8662.cif.gz | 8 KB | ||
| Others |  emd_8662_additional.map.gz emd_8662_additional.map.gz | 588.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8662 http://ftp.pdbj.org/pub/emdb/structures/EMD-8662 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8662 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8662 | HTTPS FTP |
-Related structure data
| Related structure data |  5vfoMC  8663C  8664C  8665C  8666C  8667C  8668C  5vfpC  5vfqC  5vfrC  5vfsC  5vftC  5vfuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10090 (Title: Nucleotide-Driven Triple-State Remodeling of the AAA-ATPase Channel in the Activated Human 26S Proteasome EMPIAR-10090 (Title: Nucleotide-Driven Triple-State Remodeling of the AAA-ATPase Channel in the Activated Human 26S ProteasomeData size: 2.8 TB Data #1: Motion-corrected single frame micrographs of ATP-gS-bound human proteasome [micrographs - single frame] Data #2: Single-particle stacks of classified conformations [picked particles - multiframe - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8662.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8662.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map corrected with a B-factor of -80 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.75 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Uncorrected raw map
| File | emd_8662_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Uncorrected raw map | ||||||||||||
| Projections & Slices |
| ||||||||||||
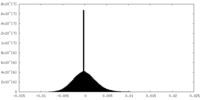
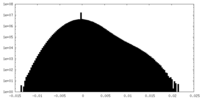
| Density Histograms |
- Sample components
Sample components
+Entire : 26S proteasome bound to ATP-gammaS
+Supramolecule #1: 26S proteasome bound to ATP-gammaS
+Macromolecule #1: Proteasome subunit alpha type-6
+Macromolecule #2: Proteasome subunit alpha type-2
+Macromolecule #3: Proteasome subunit alpha type-4
+Macromolecule #4: Proteasome subunit alpha type-7
+Macromolecule #5: Proteasome subunit alpha type-5
+Macromolecule #6: Proteasome subunit alpha type-1
+Macromolecule #7: Proteasome subunit alpha type-3
+Macromolecule #8: Proteasome subunit beta type-6
+Macromolecule #9: Proteasome subunit beta type-7
+Macromolecule #10: Proteasome subunit beta type-3
+Macromolecule #11: Proteasome subunit beta type-2
+Macromolecule #12: Proteasome subunit beta type-5
+Macromolecule #13: Proteasome subunit beta type-1
+Macromolecule #14: Proteasome subunit beta type-4
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)