+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of F-actin in the ADP state | ||||||||||||
 Map data Map data | Sharpened map of the actin filament determined by helical reconstruction. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Actin cytoskeleton / filament / F-actin / STRUCTURAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationcytoskeletal motor activator activity / tropomyosin binding / myosin heavy chain binding / mesenchyme migration / troponin I binding / filamentous actin / actin filament bundle / skeletal muscle thin filament assembly / actin filament bundle assembly / striated muscle thin filament ...cytoskeletal motor activator activity / tropomyosin binding / myosin heavy chain binding / mesenchyme migration / troponin I binding / filamentous actin / actin filament bundle / skeletal muscle thin filament assembly / actin filament bundle assembly / striated muscle thin filament / skeletal muscle myofibril / actin monomer binding / skeletal muscle fiber development / stress fiber / titin binding / actin filament polymerization / filopodium / actin filament / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / calcium-dependent protein binding / lamellipodium / cell body / hydrolase activity / protein domain specific binding / calcium ion binding / positive regulation of gene expression / magnesium ion binding / ATP binding / identical protein binding / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   | ||||||||||||
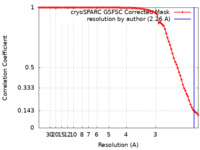
| Method | helical reconstruction / cryo EM / Resolution: 2.26 Å | ||||||||||||
 Authors Authors | Carman PJ / Barrie KR / Dominguez R | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structures of the free and capped ends of the actin filament. Authors: Peter J Carman / Kyle R Barrie / Grzegorz Rebowski / Roberto Dominguez /  Abstract: The barbed and pointed ends of the actin filament (F-actin) are the sites of growth and shrinkage and the targets of capping proteins that block subunit exchange, including CapZ at the barbed end and ...The barbed and pointed ends of the actin filament (F-actin) are the sites of growth and shrinkage and the targets of capping proteins that block subunit exchange, including CapZ at the barbed end and tropomodulin at the pointed end. We describe cryo-electron microscopy structures of the free and capped ends of F-actin. Terminal subunits at the free barbed end adopt a "flat" F-actin conformation. CapZ binds with minor changes to the barbed end but with major changes to itself. By contrast, subunits at the free pointed end adopt a "twisted" monomeric actin (G-actin) conformation. Tropomodulin binding forces the second subunit into an F-actin conformation. The structures reveal how the ends differ from the middle in F-actin and how these differences control subunit addition, dissociation, capping, and interactions with end-binding proteins. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28932.map.gz emd_28932.map.gz | 56.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28932-v30.xml emd-28932-v30.xml emd-28932.xml emd-28932.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28932_fsc.xml emd_28932_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_28932.png emd_28932.png | 975.8 KB | ||
| Others |  emd_28932_additional_1.map.gz emd_28932_additional_1.map.gz emd_28932_half_map_1.map.gz emd_28932_half_map_1.map.gz emd_28932_half_map_2.map.gz emd_28932_half_map_2.map.gz | 32.3 MB 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28932 http://ftp.pdbj.org/pub/emdb/structures/EMD-28932 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28932 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28932 | HTTPS FTP |
-Validation report
| Summary document |  emd_28932_validation.pdf.gz emd_28932_validation.pdf.gz | 733.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28932_full_validation.pdf.gz emd_28932_full_validation.pdf.gz | 733.5 KB | Display | |
| Data in XML |  emd_28932_validation.xml.gz emd_28932_validation.xml.gz | 15.8 KB | Display | |
| Data in CIF |  emd_28932_validation.cif.gz emd_28932_validation.cif.gz | 20.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28932 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28932 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28932 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28932 | HTTPS FTP |
-Related structure data
| Related structure data |  8f8pMC  8f8qC  8f8rC  8f8sC  8f8tC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28932.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28932.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of the actin filament determined by helical reconstruction. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
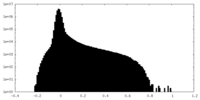
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened map of the actin filament determined by...
| File | emd_28932_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map of the actin filament determined by helical reconstruction. | ||||||||||||
| Projections & Slices |
| ||||||||||||
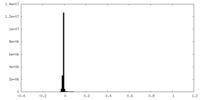
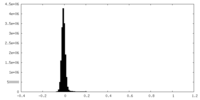
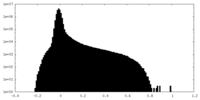
| Density Histograms |
-Half map: Half-map A of the actin filament determined by...
| File | emd_28932_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map A of the actin filament determined by helical reconstruction. | ||||||||||||
| Projections & Slices |
| ||||||||||||
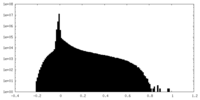
| Density Histograms |
-Half map: Half-map B of the actin filament determined by...
| File | emd_28932_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map B of the actin filament determined by helical reconstruction. | ||||||||||||
| Projections & Slices |
| ||||||||||||
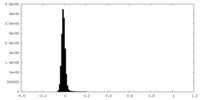
| Density Histograms |
- Sample components
Sample components
-Entire : F-actin in the ADP state
| Entire | Name: F-actin in the ADP state |
|---|---|
| Components |
|
-Supramolecule #1: F-actin in the ADP state
| Supramolecule | Name: F-actin in the ADP state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Actin, alpha skeletal muscle
| Macromolecule | Name: Actin, alpha skeletal muscle / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.109973 KDa |
| Sequence | String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIE(HIC)G IIT NWDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLD SG DGVTHNVPIY ...String: MCDEDETTAL VCDNGSGLVK AGFAGDDAPR AVFPSIVGRP RHQGVMVGMG QKDSYVGDEA QSKRGILTLK YPIE(HIC)G IIT NWDDMEKIWH HTFYNELRVA PEEHPTLLTE APLNPKANRE KMTQIMFETF NVPAMYVAIQ AVLSLYASGR TTGIVLD SG DGVTHNVPIY EGYALPHAIM RLDLAGRDLT DYLMKILTER GYSFVTTAER EIVRDIKEKL CYVALDFENE MATAASSS S LEKSYELPDG QVITIGNERF RCPETLFQPS FIGMESAGIH ETTYNSIMKC DIDIRKDLYA NNVMSGGTTM YPGIADRMQ KEITALAPST MKIKIIAPPE RKYSVWIGGS ILASLSTFQQ MWITKQEYDE AGPSIVHRKC F |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 5 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.05 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV / Details: Blot force 0 Blot time 2.5 s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 4 / Number real images: 18611 / Average electron dose: 44.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation coefficient |
|---|---|
| Output model |  PDB-8f8p: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)