[English] 日本語
 Yorodumi
Yorodumi- EMDB-27420: Structural Basis of MicroRNA Biogenesis by Dicer-1 and Its Partne... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structural Basis of MicroRNA Biogenesis by Dicer-1 and Its Partner Protein Loqs-PB - complex IV | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Dicer / Dcr-1 / Loquacious / Loqs-PB / miRNA / RNA BINDING PROTEIN-RNA complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitotic cell cycle, embryonic / germarium-derived female germ-line cyst formation / germ-line stem cell division / pole cell formation / MicroRNA (miRNA) biogenesis / segment polarity determination / Small interfering RNA (siRNA) biogenesis / germarium-derived oocyte fate determination / PKR-mediated signaling / female germ-line stem cell asymmetric division ...mitotic cell cycle, embryonic / germarium-derived female germ-line cyst formation / germ-line stem cell division / pole cell formation / MicroRNA (miRNA) biogenesis / segment polarity determination / Small interfering RNA (siRNA) biogenesis / germarium-derived oocyte fate determination / PKR-mediated signaling / female germ-line stem cell asymmetric division / regulation of regulatory ncRNA processing / RISC complex binding / dosage compensation by hyperactivation of X chromosome / pre-miRNA binding / germ-line stem cell population maintenance / ribonuclease III / apoptotic DNA fragmentation / deoxyribonuclease I activity / RISC-loading complex / miRNA metabolic process / ribonuclease III activity / RISC complex assembly / regulatory ncRNA-mediated post-transcriptional gene silencing / miRNA processing / pre-miRNA processing / siRNA processing / siRNA binding / RISC complex / dendrite morphogenesis / response to starvation / RNA endonuclease activity / helicase activity / central nervous system development / double-stranded RNA binding / single-stranded RNA binding / RNA binding / ATP binding / metal ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.73 Å | ||||||||||||
 Authors Authors | Jouravleva K / Golovenko D / Demo G / Dutcher RC / Tanaka Hall TM / Zamore PD / Korostelev AA | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: Structural basis of microRNA biogenesis by Dicer-1 and its partner protein Loqs-PB. Authors: Karina Jouravleva / Dmitrij Golovenko / Gabriel Demo / Robert C Dutcher / Traci M Tanaka Hall / Phillip D Zamore / Andrei A Korostelev /   Abstract: In animals and plants, Dicer enzymes collaborate with double-stranded RNA-binding domain (dsRBD) proteins to convert precursor-microRNAs (pre-miRNAs) into miRNA duplexes. We report six cryo-EM ...In animals and plants, Dicer enzymes collaborate with double-stranded RNA-binding domain (dsRBD) proteins to convert precursor-microRNAs (pre-miRNAs) into miRNA duplexes. We report six cryo-EM structures of Drosophila Dicer-1 that show how Dicer-1 and its partner Loqs‑PB cooperate (1) before binding pre-miRNA, (2) after binding and in a catalytically competent state, (3) after nicking one arm of the pre-miRNA, and (4) following complete dicing and initial product release. Our reconstructions suggest that pre-miRNA binds a rare, open conformation of the Dicer‑1⋅Loqs‑PB heterodimer. The Dicer-1 dsRBD and three Loqs‑PB dsRBDs form a tight belt around the pre-miRNA, distorting the RNA helix to place the scissile phosphodiester bonds in the RNase III active sites. Pre-miRNA cleavage shifts the dsRBDs and partially closes Dicer-1, which may promote product release. Our data suggest a model for how the Dicer‑1⋅Loqs‑PB complex affects a complete cycle of pre-miRNA recognition, stepwise endonuclease cleavage, and product release. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27420.map.gz emd_27420.map.gz | 294.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27420-v30.xml emd-27420-v30.xml emd-27420.xml emd-27420.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_27420.png emd_27420.png | 71.9 KB | ||
| Filedesc metadata |  emd-27420.cif.gz emd-27420.cif.gz | 7.7 KB | ||
| Others |  emd_27420_half_map_1.map.gz emd_27420_half_map_1.map.gz emd_27420_half_map_2.map.gz emd_27420_half_map_2.map.gz | 289.7 MB 289.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27420 http://ftp.pdbj.org/pub/emdb/structures/EMD-27420 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27420 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27420 | HTTPS FTP |
-Related structure data
| Related structure data |  8dgaMC  8dfvC  8dg5C  8dg7C  8dgiC  8dgjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27420.map.gz / Format: CCP4 / Size: 311.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27420.map.gz / Format: CCP4 / Size: 311.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_27420_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27420_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of Dicer-1 and Loqs-PB binding miRNA
| Entire | Name: Ternary complex of Dicer-1 and Loqs-PB binding miRNA |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of Dicer-1 and Loqs-PB binding miRNA
| Supramolecule | Name: Ternary complex of Dicer-1 and Loqs-PB binding miRNA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Endoribonuclease Dcr-1
| Macromolecule | Name: Endoribonuclease Dcr-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on ester bonds; Endoribonucleases producing 5'-phosphomonoesters |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 255.733797 KDa |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Sequence | String: MAFHWCDNNL HTTVFTPRDF QVELLATAYE RNTIICLGHR SSKEFIALKL LQELSRRARR HGRVSVYLSC EVGTSTEPCS IYTMLTHLT DLRVWQEQPD MQIPFDHCWT DYHVSILRPE GFLYLLETRE LLLSRVELIV LEDCHDSAVY QRIRPLFENH I MPAPPADR ...String: MAFHWCDNNL HTTVFTPRDF QVELLATAYE RNTIICLGHR SSKEFIALKL LQELSRRARR HGRVSVYLSC EVGTSTEPCS IYTMLTHLT DLRVWQEQPD MQIPFDHCWT DYHVSILRPE GFLYLLETRE LLLSRVELIV LEDCHDSAVY QRIRPLFENH I MPAPPADR PRILGLAGPL HSAGCELQQL SAMLATLEQS VLCQIESASD IVTVLRYCSR PHEYIVQCAP FEMDELSLVL AD VLNTHKS FLLDHRYDPY EIYGTDQFMD ELKDIPDPKV DPLNVINSLL VVLHEMGPWC TQRAAHHFYQ CNEKLKVKTP HER HYLLYC LVSTALIQLY SLCEHAFHRH LGSGSDSRQT IERYSSPKVR RLLQTLRCFK PEEVHTQADG LRRMRHQVDQ ADFN RLSHT LESKCRMVDQ LDQPPTETRA LVATLEQILH TTEDRQTNRS AARVTPTPTP AHAKPKPSSG ANTAQPRTRR RVYTR RHHR DHNDGSDTLC ALIYCNQNHT ARVLFELLAE ISRRDPDLKF LRCQYTTDRV ADPTTEPKEA ELEHRRQEEV LKRFRM HDC NVLIGTSVLE EGIDVPKCNL VVRWDPPTTY RSYVQCKGRA RAAPAYHVIL VAPSYKSPTV GSVQLTDRSH RYICATG DT TEADSDSDDS AMPNSSGSDP YTFGTARGTV KILNPEVFSK QPPTACDIKL QEIQDELPAS AQLDTSNSSD EAVSMSNT S PSESSTEQKS RRFQCELSSL TEPEDTSDTT AEIDTAHSLA STTKDLVHQM AQYREIEQML LSKCANTEPP EQEQCEAER FSACLAAYRP KPHLLTGASV DLGSAIALVN KYCARLPSDT FTKLTALWRC TRNERAGVTL FQYTLRLPIN SPLKHDIVGL PMPTQTLAR RLAALQACVE LHRIGELDDQ LQPIGKEGFR ALEPDWECFE LEPEDEQIVQ LSDEPRPGTT KRRQYYYKRI A SEFCDCRP VAGAPCYLYF IQLTLQCPIP EEQNTRGRKI YPPEDAQQGF GILTTKRIPK LSAFSIFTRS GEVKVSLELA KE RVILTSE QIVCINGFLN YTFTNVLRLQ KFLMLFDPDS TENCVFIVPT VKAPAGGKHI DWQFLELIQA NGNTMPRAVP DEE RQAQPF DPQRFQDAVV MPWYRNQDQP QYFYVAEICP HLSPLSCFPG DNYRTFKHYY LVKYGLTIQN TSQPLLDVDH TSAR LNFLT PRYVNRKGVA LPTSSEETKR AKRENLEQKQ ILVPELCTVH PFPASLWRTA VCLPCILYRI NGLLLADDIR KQVSA DLGL GRQQIEDEDF EWPMLDFGWS LSEVLKKSRE SKQKESLKDD TINGKDLVDV EKKAISEETQ IDKDSKDDKV EKSAIE LII EGEEKLQEAD DFIEIGTWSN DMADDIASFN QEDDDEDDAF HLPVLPANVK FCDQQTRYGS PTFWDVSNGE SGFKGPK SS QNKQGGKGKA KGPAKPTFNY YDSDNSLGSS YDDDDNAGPL NYMHHNYSSD DDDVADDIDA GRIAFTSKNE AETIETAQ E VEKRQKQLSI IQATNANERQ YQQTKNLLIG FNFKHEDQKE PATIRYEESI AKLKTEIESG GMLVPHDQQL VLKRSDAAE AQVAKVSMME LLKQLLPYVN EDVLAKKLGD RRELLLSDLV ELNADWVARH EQETYNVMGC GDSFDNYNDH HRLNLDEKQL KLQYERIEI EPPTSTKAIT SAILPAGFSF DRQPDLVGHP GPSPSIILQA LTMSNANDGI NLERLETIGD SFLKYAITTY L YITYENVH EGKLSHLRSK QVANLNLYRL GRRKRLGEYM IATKFEPHDN WLPPCYYVPK ELEKALIEAK IPTHHWKLAD LL DIKNLSS VQICEMVREK ADALGLEQNG GAQNGQLDDS NDSCNDFSCF IPYNLVSQHS IPDKSIADCV EALIGAYLIE CGP RGALLF MAWLGVRVLP ITRQLDGGNQ EQRIPGSTKP NAENVVTVYG AWPTPRSPLL HFAPNATEEL DQLLSGFEEF EESL GYKFR DRSYLLQAMT HASYTPNRLT DCYQRLEFLG DAVLDYLITR HLYEDPRQHS PGALTDLRSA LVNNTIFASL AVRHG FHKF FRHLSPGLND VIDRFVRIQQ ENGHCISEEY YLLSEEECDD AEDVEVPKAL GDVFESIAGA IFLDSNMSLD VVWHVY SNM MSPEIEQFSN SVPKSPIREL LELEPETAKF GKPEKLADGR RVRVTVDVFC KGTFRGIGRN YRIAKCTAAK CALRQLK KQ GLIAKKD UniProtKB: Endoribonuclease Dcr-1 |
-Macromolecule #4: Loquacious, isoform B
| Macromolecule | Name: Loquacious, isoform B / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 50.149867 KDa |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Sequence | String: MDQENFHGSS LPQQLQNLHI QPQQASPNPV QTGFAPRRHY NNLVGLGNGN AVSGSPVKGA PLGQRHVKLK KEKISAQVAQ LSQPGQLQL SDVGDPALAG GSGLQGGVGL MGVILPSDEA LKFVSETDAN GLAMKTPVSI LQELLSRRGI TPGYELVQIE G AIHEPTFR ...String: MDQENFHGSS LPQQLQNLHI QPQQASPNPV QTGFAPRRHY NNLVGLGNGN AVSGSPVKGA PLGQRHVKLK KEKISAQVAQ LSQPGQLQL SDVGDPALAG GSGLQGGVGL MGVILPSDEA LKFVSETDAN GLAMKTPVSI LQELLSRRGI TPGYELVQIE G AIHEPTFR FRVSFKDKDT PFTAMGAGRS KKEAKHAAAR ALIDKLIGAQ LPESPSSSAG PSVTGLTVAG SGGDGNANAT GG GDASDKT VGNPIGWLQE MCMQRRWPPP SYETETEVGL PHERLFTIAC SILNYREMGK GKSKKIAKRL AAHRMWMRLQ ETP IDSGKI SDSICGELEG EPRSSENYYG ELKDISVPTL TTQHSNKVSQ FHKTLKNATG KKLLKLQKTC LKNNKIDYIK LLGE IATEN QFEVTYVDIE EKTFSGQFQC LVQLSTLPVG VCHGSGPTAA DAQRHAAQNA LEYLKIMTKK UniProtKB: Protein Loquacious |
-Macromolecule #2: RNA (22-MER)
| Macromolecule | Name: RNA (22-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.835085 KDa |
| Sequence | String: GAGGUAGUAG GUUGUAUAGU A |
-Macromolecule #3: RNA (22-MER)
| Macromolecule | Name: RNA (22-MER) / type: rna / ID: 3 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.820074 KDa |
| Sequence | String: CUAUACAACC UACUACCUCU CU |
-Macromolecule #5: URIDINE-5'-MONOPHOSPHATE
| Macromolecule | Name: URIDINE-5'-MONOPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: U5P |
|---|---|
| Molecular weight | Theoretical: 324.181 Da |
| Chemical component information | 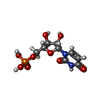 ChemComp-U: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.9 / Component - Concentration: 20.0 mM / Component - Formula: HEPES-KOH / Component - Name: HEPES-KOH |
| Grid | Model: C-flat-1.2/1.3 / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 66.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)