[English] 日本語
 Yorodumi
Yorodumi- EMDB-15284: Cryo-EM structure of USP1-UAF1 bound to FANCI and mono-ubiquitina... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of USP1-UAF1 bound to FANCI and mono-ubiquitinated FANCD2 without ML323 (consensus reconstruction) | |||||||||
 Map data Map data | Globally sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Deubiquitinase / Complex / Enzyme-Substrate / Inhibitor / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of protein monoubiquitination / positive regulation of error-prone translesion synthesis / Signaling by cytosolic PDGFRA and PDGFRB fusion proteins / regulation of CD40 signaling pathway / gamete generation / monoubiquitinated protein deubiquitination / regulation of regulatory T cell differentiation / homologous chromosome pairing at meiosis / neuronal stem cell population maintenance / double-strand break repair involved in meiotic recombination ...regulation of protein monoubiquitination / positive regulation of error-prone translesion synthesis / Signaling by cytosolic PDGFRA and PDGFRB fusion proteins / regulation of CD40 signaling pathway / gamete generation / monoubiquitinated protein deubiquitination / regulation of regulatory T cell differentiation / homologous chromosome pairing at meiosis / neuronal stem cell population maintenance / double-strand break repair involved in meiotic recombination / brain morphogenesis / deubiquitinase activator activity / DNA repair complex / mitotic intra-S DNA damage checkpoint signaling / skeletal system morphogenesis / skin development / seminiferous tubule development / homeostasis of number of cells / embryonic organ development / protein deubiquitination / single fertilization / regulation of DNA repair / response to UV / interstrand cross-link repair / positive regulation of double-strand break repair via homologous recombination / condensed chromosome / DNA polymerase binding / Maturation of protein E / Maturation of protein E / ER Quality Control Compartment (ERQC) / Myoclonic epilepsy of Lafora / FLT3 signaling by CBL mutants / IRAK2 mediated activation of TAK1 complex / Alpha-protein kinase 1 signaling pathway / Glycogen synthesis / IRAK1 recruits IKK complex / IRAK1 recruits IKK complex upon TLR7/8 or 9 stimulation / Prevention of phagosomal-lysosomal fusion / Endosomal Sorting Complex Required For Transport (ESCRT) / Membrane binding and targetting of GAG proteins / Negative regulation of FLT3 / Regulation of TBK1, IKKε (IKBKE)-mediated activation of IRF3, IRF7 / PTK6 Regulates RTKs and Their Effectors AKT1 and DOK1 / Regulation of TBK1, IKKε-mediated activation of IRF3, IRF7 upon TLR3 ligation / IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation / Constitutive Signaling by NOTCH1 HD Domain Mutants / NOTCH2 Activation and Transmission of Signal to the Nucleus / TICAM1,TRAF6-dependent induction of TAK1 complex / TICAM1-dependent activation of IRF3/IRF7 / APC/C:Cdc20 mediated degradation of Cyclin B / Downregulation of ERBB4 signaling / Regulation of FZD by ubiquitination / APC-Cdc20 mediated degradation of Nek2A / p75NTR recruits signalling complexes / InlA-mediated entry of Listeria monocytogenes into host cells / TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling / NF-kB is activated and signals survival / TRAF6-mediated induction of TAK1 complex within TLR4 complex / Regulation of pyruvate metabolism / Pexophagy / Downregulation of ERBB2:ERBB3 signaling / Regulation of innate immune responses to cytosolic DNA / NRIF signals cell death from the nucleus / Regulation of PTEN localization / positive regulation of epithelial cell proliferation / VLDLR internalisation and degradation / positive regulation of protein ubiquitination / Activated NOTCH1 Transmits Signal to the Nucleus / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / Translesion synthesis by REV1 / TICAM1, RIP1-mediated IKK complex recruitment / Regulation of BACH1 activity / Translesion synthesis by POLK / InlB-mediated entry of Listeria monocytogenes into host cell / JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 / MAP3K8 (TPL2)-dependent MAPK1/3 activation / Activation of IRF3, IRF7 mediated by TBK1, IKKε (IKBKE) / Downregulation of TGF-beta receptor signaling / Translesion synthesis by POLI / Josephin domain DUBs / Gap-filling DNA repair synthesis and ligation in GG-NER / IKK complex recruitment mediated by RIP1 / PINK1-PRKN Mediated Mitophagy / response to gamma radiation / TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) / Regulation of activated PAK-2p34 by proteasome mediated degradation / TNFR1-induced NF-kappa-B signaling pathway / positive regulation of receptor signaling pathway via JAK-STAT / TCF dependent signaling in response to WNT / skeletal system development / Regulation of NF-kappa B signaling / activated TAK1 mediates p38 MAPK activation / TP53 Regulates Transcription of DNA Repair Genes / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / ubiquitin binding / Asymmetric localization of PCP proteins / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / Negative regulators of DDX58/IFIH1 signaling / Ubiquitin-dependent degradation of Cyclin D Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||
 Authors Authors | Rennie ML / Walden H | |||||||||
| Funding support | European Union,  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Cryo-EM reveals a mechanism of USP1 inhibition through a cryptic binding site. Authors: Martin L Rennie / Connor Arkinson / Viduth K Chaugule / Helen Walden /  Abstract: Repair of DNA damage is critical to genomic integrity and frequently disrupted in cancers. Ubiquitin-specific protease 1 (USP1), a nucleus-localized deubiquitinase, lies at the interface of multiple ...Repair of DNA damage is critical to genomic integrity and frequently disrupted in cancers. Ubiquitin-specific protease 1 (USP1), a nucleus-localized deubiquitinase, lies at the interface of multiple DNA repair pathways and is a promising drug target for certain cancers. Although multiple inhibitors of this enzyme, including one in phase 1 clinical trials, have been established, their binding mode is unknown. Here, we use cryo-electron microscopy to study an assembled enzyme-substrate-inhibitor complex of USP1 and the well-established inhibitor, ML323. Achieving 2.5-Å resolution, with and without ML323, we find an unusual binding mode in which the inhibitor disrupts part of the hydrophobic core of USP1. The consequent conformational changes in the secondary structure lead to subtle rearrangements in the active site that underlie the mechanism of inhibition. These structures provide a platform for structure-based drug design targeting USP1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15284.map.gz emd_15284.map.gz | 118.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15284-v30.xml emd-15284-v30.xml emd-15284.xml emd-15284.xml | 28.7 KB 28.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15284_fsc.xml emd_15284_fsc.xml | 11.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_15284.png emd_15284.png | 181.5 KB | ||
| Masks |  emd_15284_msk_1.map emd_15284_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15284.cif.gz emd-15284.cif.gz | 9.2 KB | ||
| Others |  emd_15284_half_map_1.map.gz emd_15284_half_map_1.map.gz emd_15284_half_map_2.map.gz emd_15284_half_map_2.map.gz | 116 MB 116 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15284 http://ftp.pdbj.org/pub/emdb/structures/EMD-15284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15284 | HTTPS FTP |
-Related structure data
| Related structure data |  8a9jMC  7zh3C  7zh4C  8a9kC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_15284.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15284.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Globally sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15284_msk_1.map emd_15284_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
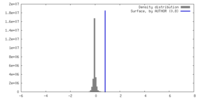
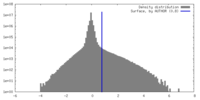
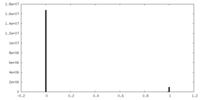
| Density Histograms |
-Half map: #1
| File | emd_15284_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_15284_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : USP1(C90S)-UAF1 bound to FANCI and mono-ubiquitinated FANCD2 with...
| Entire | Name: USP1(C90S)-UAF1 bound to FANCI and mono-ubiquitinated FANCD2 with dsDNA |
|---|---|
| Components |
|
-Supramolecule #1: USP1(C90S)-UAF1 bound to FANCI and mono-ubiquitinated FANCD2 with...
| Supramolecule | Name: USP1(C90S)-UAF1 bound to FANCI and mono-ubiquitinated FANCD2 with dsDNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|
-Supramolecule #2: Fanconi anemia group I protein, Fanconi anemia group D2 protein w...
| Supramolecule | Name: Fanconi anemia group I protein, Fanconi anemia group D2 protein with ubiquitin conjugated to K561, Ubiquitin carboxyl-terminal hydrolase 1, WD repeat-containing protein 48 type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Fanconi anemia group I protein
| Macromolecule | Name: Fanconi anemia group I protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150.459125 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHMDQ KILSLAAEKT ADKLQEFLQT LREGDLTNLL QNQAVKGKVA GALLRAIFKG SPCSEEAGTL RRRKIYTCCI QLVESGDLQ KEIASEIIGL LMLEAHHFPG PLLVELANEF ISAVREGSLV NGKSLELLPI ILTALATKKE NLAYGKGVLS G EECKKQLI ...String: MHHHHHHMDQ KILSLAAEKT ADKLQEFLQT LREGDLTNLL QNQAVKGKVA GALLRAIFKG SPCSEEAGTL RRRKIYTCCI QLVESGDLQ KEIASEIIGL LMLEAHHFPG PLLVELANEF ISAVREGSLV NGKSLELLPI ILTALATKKE NLAYGKGVLS G EECKKQLI NTLCSGRWDQ QYVIQLTSMF KDVPLTAEEV EFVVEKALSM FSKMNLQEIP PLVYQLLVLS SKGSRKSVLE GI IAFFSAL DKQHNEEQSG DELLDVVTVP SGELRHVEGT IILHIVFAIK LDYELGRELV KHLKVGQQGD SNNNLSPFSI ALL LSVTRI QRFQDQVLDL LKTSVVKSFK DLQLLQGSKF LQNLVPHRSY VSTMILEVVK NSVHSWDHVT QGLVELGFIL MDSY GPKKV LDGKTIETSP SLSRMPNQHA CKLGANILLE TFKIHEMIRQ EILEQVLNRV VTRASSPISH FLDLLSNIVM YAPLV LQSC SSKVTEAFDY LSFLPLQTVQ RLLKAVQPLL KVSMSMRDCL ILVLRKAMFA NQLDARKSAV AGFLLLLKNF KVLGSL SSS QCSQSLSVSQ VHVDVHSHYN SVANETFCLE IMDSLRRCLS QQADVRLMLY EGFYDVLRRN SQLANSVMQT LLSQLKQ FY EPKPDLLPPL KLEACILTQG DKISLQEPLD YLLCCIQHCL AWYKNTVIPL QQGEEEEEEE EAFYEDLDDI LESITNRM I KSELEDFELD KSADFSQSTS IGIKNNICAF LVMGVCEVLI EYNFSISSFS KNRFEDILSL FMCYKKLSDI LNEKAGKAK TKMANKTSDS LLSMKFVSSL LTALFRDSIQ SHQESLSVLR SSNEFMRYAV NVALQKVQQL KETGHVSGPD GQNPEKIFQN LCDITRVLL WRYTSIPTSV EESGKKEKGK SISLLCLEGL QKIFSAVQQF YQPKIQQFLR ALDVTDKEGE EREDADVSVT Q RTAFQIRQ FQRSLLNLLS SQEEDFNSKE ALLLVTVLTS LSKLLEPSSP QFVQMLSWTS KICKENSRED ALFCKSLMNL LF SLHVSYK SPVILLRDLS QDIHGHLGDI DQDVEVEKTN HFAIVNLRTA APTVCLLVLS QAEKVLEEVD WLITKLKGQV SQE TLSEEA SSQATLPNQP VEKAIIMQLG TLLTFFHELV QTALPSGSCV DTLLKDLCKM YTTLTALVRY YLQVCQSSGG IPKN MEKLV KLSGSHLTPL CYSFISYVQN KSKSLNYTGE KKEKPAAVAT AMARVLRETK PIPNLIFAIE QYEKFLIHLS KKSKV NLMQ HMKLSTSRDF KIKGNILDMV LREDGEDENE EGTASEHGGQ NKEPAKKKRK K UniProtKB: Fanconi anemia group I protein |
-Macromolecule #2: Fanconi anemia group D2 protein
| Macromolecule | Name: Fanconi anemia group D2 protein / type: protein_or_peptide / ID: 2 / Details: Ubiquitin conjugated to K561 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 164.623828 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGSMVSKRR LSKSEDKESL TEDASKTRKQ PLSKKTKKSH IANEVEENDS IFVKLLKISG IILKTGESQN QLAVDQIAFQ KKLFQTLRR HPSYPKIIEE FVSGLESYIE DEDSFRNCLL SCERLQDEEA SMGASYSKSL IKLLLGIDIL QPAIIKTLFE K LPEYFFEN ...String: GPGSMVSKRR LSKSEDKESL TEDASKTRKQ PLSKKTKKSH IANEVEENDS IFVKLLKISG IILKTGESQN QLAVDQIAFQ KKLFQTLRR HPSYPKIIEE FVSGLESYIE DEDSFRNCLL SCERLQDEEA SMGASYSKSL IKLLLGIDIL QPAIIKTLFE K LPEYFFEN KNSDEINIPR LIVSQLKWLD RVVDGKDLTT KIMQLISIAP ENLQHDIITS LPEILGDSQH ADVGKELSDL LI ENTSLTV PILDVLSSLR LDPNFLLKVR QLVMDKLSSI RLEDLPVIIK FILHSVTAMD TLEVISELRE KLDLQHCVLP SRL QASQVK LKSKGRASSS GNQESSGQSC IILLFDVIKS AIRYEKTISE AWIKAIENTA SVSEHKVFDL VMLFIIYSTN TQTK KYIDR VLRNKIRSGC IQEQLLQSTF SVHYLVLKDM CSSILSLAQS LLHSLDQSII SFGSLLYKYA FKFFDTYCQQ EVVGA LVTH ICSGNEAEVD TALDVLLELV VLNPSAMMMN AVFVKGILDY LDNISPQQIR KLFYVLSTLA FSKQNEASSH IQDDMH LVI RKQLSSTVFK YKLIGIIGAV TMAGIMAADR SESPSLTQER ANLSDEQCTQ VTSLLQLVHS CSEQSPQASA LYYDEFA NL IQHEKLDPKA LEWVGHTICN DFQDAFVVDS CVVPEGDFPF PVKALYGLEE YDTQDGIAIN LLPLLFSQDF AKDGGPVT S QESGQKLVSP LCLAPYFRLL RLCVERQHNG NLEEIDGLLD CPIFLTDLEP GEKLESMSAK ERSFMCSLIF LTLNWFREI VNAFCQETSP EMKGKVLTRL KHIVELQIIL EKYLAVTPDY VPPLGNFDVE TLDITPHTVT AISAKIRKKG KIERKQKTDG SKTSSSDTL SEEKNSECDP TPSHRGQLNK EFTGKEEKTS LLLHNSHAFF RELDIEVFSI LHCGLVTKFI LDTEMHTEAT E VVQLGPPE LLFLLEDLSQ KLESMLTPPI ARRVPFLKNK GSRNIGFSHL QQRSAQEIVH CVFQLLTPMC NHLENIHNYF QC LAAENHG VVDGPGVKVQ EYHIMSSCYQ RLLQIFHGLF AWSGFSQPEN QNLLYSALHV LSSRLKQGEH SQPLEELLSQ SVH YLQNFH QSIPSFQCAL YLIRLLMVIL EKSTASAQNK EKIASLARQF LCRVWPSGDK EKSNISNDQL HALLCIYLEH TESI LKAIE EIAGVGVPEL INSPKDASSS TFPTLTRHTF VVFFRVMMAE LEKTVKKIEP GTAADSQQIH EEKLLYWNMA VRDFS ILIN LIKVFDSHPV LHVCLKYGRL FVEAFLKQCM PLLDFSFRKH REDVLSLLET FQLDTRLLHH LCGHSKIHQD TRLTQH VPL LKKTLELLVC RVKAMLTLNN CREAFWLGNL KNRDLQGEEI KSQNSQESTA DESEDDMSSQ ASKSKATEDG EEDEVSA GE KEQDSDESYD DSD UniProtKB: Fanconi anemia group D2 protein |
-Macromolecule #3: Polyubiquitin-C
| Macromolecule | Name: Polyubiquitin-C / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.875125 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGSMQIFVK TLTGKTITLE VEPSDTIENV KAKIQDKEGI PPDQQRLIFA GKQLEDGRTL SDYNIQKEST LHLVLRLRGG UniProtKB: Polyubiquitin-C |
-Macromolecule #4: Ubiquitin carboxyl-terminal hydrolase 1
| Macromolecule | Name: Ubiquitin carboxyl-terminal hydrolase 1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: ubiquitinyl hydrolase 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 88.390273 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GMPGVIPSES NGLSRGSPSK KNRLSLKFFQ KKETKRALDF TDSQENEEKA SEYRASEIDQ VVPAAQSSPI NCEKRENLLP FVGLNNLGN TSYLNSILQV LYFCPGFKSG VKHLFNIISR KKEALKDEAN QKDKGNCKED SLASYELICS LQSLIISVEQ L QASFLLNP ...String: GMPGVIPSES NGLSRGSPSK KNRLSLKFFQ KKETKRALDF TDSQENEEKA SEYRASEIDQ VVPAAQSSPI NCEKRENLLP FVGLNNLGN TSYLNSILQV LYFCPGFKSG VKHLFNIISR KKEALKDEAN QKDKGNCKED SLASYELICS LQSLIISVEQ L QASFLLNP EKYTDELATQ PRRLLNTLRE LNPMYEGYLQ HDAQEVLQCI LGNIQETCQL LKKEEVKNVA ELPTKVEEIP HP KEEMNGI NSIEMDSMRH SEDFKEKLPK GNGKRKSDTE FGNMKKKVKL SKEHQSLEEN QRQTRSKRKA TSDTLESPPK IIP KYISEN ESPRPSQKKS RVKINWLKSA TKQPSILSKF CSLGKITTNQ GVKGQSKENE CDPEEDLGKC ESDNTTNGCG LESP GNTVT PVNVNEVKPI NKGEEQIGFE LVEKLFQGQL VLRTRCLECE SLTERREDFQ DISVPVQEDE LSKVEESSEI SPEPK TEMK TLRWAISQFA SVERIVGEDK YFCENCHHYT EAERSLLFDK MPEVITIHLK CFAASGLEFD CYGGGLSKIN TPLLTP LKL SLEEWSTKPT NDSYGLFAVV MHSGITISSG HYTASVKVTD LNSLELDKGN FVVDQMCEIG KPEPLNEEEA RGVVENY ND EEVSIRVGGN TQPSKVLNKK NVEAIGLLAA QKSKADYELY NKASNPDKVA STAFAENRNS ETSDTTGTHE SDRNKESS D QTGINISGFE NKISYVVQSL KEYEGKWLLF DDSEVKVTEE KDFLNSLSPS TSPTSTPYLL FYKKL UniProtKB: Ubiquitin carboxyl-terminal hydrolase 1 |
-Macromolecule #5: WD repeat-containing protein 48
| Macromolecule | Name: WD repeat-containing protein 48 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 78.300656 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHLEV LFQGPGSMAA HHRQNTAGRR KVQVSYVIRD EVEKYNRNGV NALQLDPALN RLFTAGRDSI IRIWSVNQHK QDPYIASME HHTDWVNDIV LCCNGKTLIS ASSDTTVKVW NAHKGFCMST LRTHKDYVKA LAYAKDKELV ASAGLDRQIF L WDVNTLTA ...String: MHHHHHHLEV LFQGPGSMAA HHRQNTAGRR KVQVSYVIRD EVEKYNRNGV NALQLDPALN RLFTAGRDSI IRIWSVNQHK QDPYIASME HHTDWVNDIV LCCNGKTLIS ASSDTTVKVW NAHKGFCMST LRTHKDYVKA LAYAKDKELV ASAGLDRQIF L WDVNTLTA LTASNNTVTT SSLSGNKDSI YSLAMNQLGT IIVSGSTEKV LRVWDPRTCA KLMKLKGHTD NVKALLLNRD GT QCLSGSS DGTIRLWSLG QQRCIATYRV HDEGVWALQV NDAFTHVYSG GRDRKIYCTD LRNPDIRVLI CEEKAPVLKM ELD RSADPP PAIWVATTKS TVNKWTLKGI HNFRASGDYD NDCTNPITPL CTQPDQVIKG GASIIQCHIL NDKRHILTKD TNNN VAYWD VLKACKVEDL GKVDFEDEIK KRFKMVYVPN WFSVDLKTGM LTITLDESDC FAAWVSAKDA GFSSPDGSDP KLNLG GLLL QALLEYWPRT HVNPMDEEEN EVNHVNGEQE NRVQKGNGYF QVPPHTPVIF GEAGGRTLFR LLCRDSGGET ESMLLN ETV PQWVIDITVD KNMPKFNKIP FYLQPHASSG AKTLKKDRLS ASDMLQVRKV MEHVYEKIIN LDNESQTTSS SNNEKPG EQ EKEEDIAVLA EEKIELLCQD QVLDPNMDLR TVKHFIWKSG GDLTLHYRQK ST UniProtKB: WD repeat-containing protein 48 |
-Macromolecule #6: DNA (61-MER)
| Macromolecule | Name: DNA (61-MER) / type: dna / ID: 6 Details: Actual sequence: TGATCAGAGGTCATTTGAATTCATGGCTTCGAGCTTCATGTAGAGTCGACGGTGCTGGGAT Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 5.17782 KDa |
| Sequence | String: (DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN) (DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN) (DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN)(DN) |
-Macromolecule #7: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 7 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 288 K / Details: blotted for 3.0 secs before plunging. |
| Details | 9.3 uM USP1-UAF1, 1.8 uM FANCI-FANCD2Ub, 2.2 uM dsDNA (61 base-pairs), 18 uM ML323 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Overall B value: 94.9 | ||||||||||||||
| Output model |  PDB-8a9j: |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)