+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tail tip of siphophage T5 : central fibre | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacteriophage / Siphophage / T5 / baseplate / VIRAL PROTEIN / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |  Escherichia phage T5 (virus) / Escherichia phage T5 (virus) /  Escherichia virus T5 Escherichia virus T5 | |||||||||
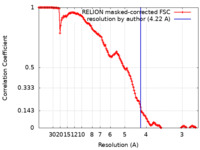
| Method | single particle reconstruction / cryo EM / Resolution: 4.22 Å | |||||||||
 Authors Authors | Linares R / Arnaud CA / Effantin G / Epalle N / Boeri Erba E / Schoehn G / Breyton C | |||||||||
| Funding support |  France, 2 items France, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2023 Journal: Sci Adv / Year: 2023Title: Structural basis of bacteriophage T5 infection trigger and cell wall perforation. Authors: Romain Linares / Charles-Adrien Arnaud / Grégory Effantin / Claudine Darnault / Nathan Hugo Epalle / Elisabetta Boeri Erba / Guy Schoehn / Cécile Breyton /  Abstract: Most bacteriophages present a tail allowing host recognition, cell wall perforation, and viral DNA channeling from the capsid to the infected bacterium cytoplasm. The majority of tailed phages bear a ...Most bacteriophages present a tail allowing host recognition, cell wall perforation, and viral DNA channeling from the capsid to the infected bacterium cytoplasm. The majority of tailed phages bear a long flexible tail () at the tip of which receptor binding proteins (RBPs) specifically interact with their host, triggering infection. In siphophage T5, the unique RBP is located at the extremity of a central fiber. We present the structures of T5 tail tip, determined by cryo-electron microscopy before and after interaction with its receptor, FhuA, reconstituted into nanodisc. These structures bring out the important conformational changes undergone by T5 tail tip upon infection, which include bending of T5 central fiber on the side of the tail tip, tail anchoring to the membrane, tail tube opening, and formation of a transmembrane channel. The data allow to detail the first steps of an otherwise undescribed infection mechanism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14790.map.gz emd_14790.map.gz | 15 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14790-v30.xml emd-14790-v30.xml emd-14790.xml emd-14790.xml | 20.6 KB 20.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14790_fsc.xml emd_14790_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_14790.png emd_14790.png | 43.5 KB | ||
| Masks |  emd_14790_msk_1.map emd_14790_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14790.cif.gz emd-14790.cif.gz | 6.4 KB | ||
| Others |  emd_14790_additional_1.map.gz emd_14790_additional_1.map.gz emd_14790_half_map_1.map.gz emd_14790_half_map_1.map.gz emd_14790_half_map_2.map.gz emd_14790_half_map_2.map.gz | 194.3 MB 194 MB 194.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14790 http://ftp.pdbj.org/pub/emdb/structures/EMD-14790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14790 | HTTPS FTP |
-Related structure data
| Related structure data |  7zlvMC  7qg9C  7zhjC  7zn2C  7zn4C  7zqbC  7zqpC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14790.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14790.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.351 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14790_msk_1.map emd_14790_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: unsharpened map
| File | emd_14790_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14790_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_14790_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Escherichia virus T5
| Entire | Name:  Escherichia virus T5 Escherichia virus T5 |
|---|---|
| Components |
|
-Supramolecule #1: Escherichia virus T5
| Supramolecule | Name: Escherichia virus T5 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Pure T5 tails obtained by infecting E. coli F strain with the amber mutant phage T5D20am30d, incubated with E. coli receptor FhuA reconstituted into nanodisc NCBI-ID: 2695836 / Sci species name: Escherichia virus T5 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
-Macromolecule #1: Probable central straight fiber
| Macromolecule | Name: Probable central straight fiber / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage T5 (virus) Escherichia phage T5 (virus) |
| Molecular weight | Theoretical: 74.851703 KDa |
| Sequence | String: MISNNAPAKM VLNSVLTGYT LAYIQHSIYS DYDVIGRSFW LKEGSNVTRR DFTGIDTFSV TINNLKPTTT YEVQGAFYDS IIDSELLNA QIGINLSDKQ TFKMKSAPRI TGARCESEPV DVGVGAPIVY IDTTGEADYC TIELKDNSNA NNPWVKYYVG A LMPTIMFG ...String: MISNNAPAKM VLNSVLTGYT LAYIQHSIYS DYDVIGRSFW LKEGSNVTRR DFTGIDTFSV TINNLKPTTT YEVQGAFYDS IIDSELLNA QIGINLSDKQ TFKMKSAPRI TGARCESEPV DVGVGAPIVY IDTTGEADYC TIELKDNSNA NNPWVKYYVG A LMPTIMFG GVPIGSYKVR ISGQISLPDG VTIDSSGYYE YPNVFEVRYN FVPPAAPINI VFKAARIADG KERYDLRVQW DW NRGAGAN VREFVLSYID SAEFVRTGWT KAQKINVGAA QSATIISFPW KVEHKFKVSS IAWGPDAQDV TDSAVQTFIL NES TPLDNS FVNETGIEVN YAYIKGKIKD GSTWKQTFLI DAATGAINIG LLDAEGKAPI SFDPVKKIVN VDGSVITKTI NAAN FVMTN LTGQDNPAIY TQGKTWGDTK SGIWMGMDNV TAKPKLDIGN ATQYIRYDGN ILRISSEVVI GTPNGDIDIQ TGIQG KQTV FIYIIGTSLP AKPTSPAYPP SGWSKTPPNR TSNTQNIYCS TGTLDPVTNQ LVSGTSWSDV VQWSGTEGVD GRPGAT GQR GPGMYSLAIA NLTAWNDSQA NSFFTSNFGS GPVKYDVLTE YKSGAPGTAF TRQWNGSAWT SPAMVLHGDM IVNGTVT AS KIVANNAFLS QIGVNIIYDR AAALSSNPEG SYKMKIDLQN GYIHIR UniProtKB: Straight fiber protein pb4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER/RHODIUM / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Details: Intensity 25 mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K / Instrument: FEI VITROBOT MARK IV Details: 3 uL of T5 tails sample (with or without FhuA-ND) were deposited on a freshly glow discharged EM grid and plunge-frozen in nitrogen-cooled liquid ethane using a ThermoFisher Mark IV Vitrobot ...Details: 3 uL of T5 tails sample (with or without FhuA-ND) were deposited on a freshly glow discharged EM grid and plunge-frozen in nitrogen-cooled liquid ethane using a ThermoFisher Mark IV Vitrobot device (100 percent humidity, 20 Celsius degrees, 5 s blotting time, blot force 0). | |||||||||||||||
| Details | Pure T5 tails obtained by infecting E. coli F strain with the amber mutant phage T5D20am30d, incubated with E. coli receptor FhuA reconstituted into nanodisc |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7zlv: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)