[English] 日本語
 Yorodumi
Yorodumi- EMDB-12931: Mouse RNF213:UBE2L3 transthiolation intermediate, chemically stab... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mouse RNF213:UBE2L3 transthiolation intermediate, chemically stabilized | |||||||||||||||
 Map data Map data | full reconstructed density | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationlipid ubiquitination / lipid droplet formation / negative regulation of non-canonical Wnt signaling pathway / cell cycle phase transition / ubiquitin-protein transferase activator activity / xenophagy / Antigen processing: Ubiquitination & Proteasome degradation / protein K11-linked ubiquitination / Transferases; Acyltransferases; Aminoacyltransferases / cellular response to glucocorticoid stimulus ...lipid ubiquitination / lipid droplet formation / negative regulation of non-canonical Wnt signaling pathway / cell cycle phase transition / ubiquitin-protein transferase activator activity / xenophagy / Antigen processing: Ubiquitination & Proteasome degradation / protein K11-linked ubiquitination / Transferases; Acyltransferases; Aminoacyltransferases / cellular response to glucocorticoid stimulus / sprouting angiogenesis / positive regulation of ubiquitin-protein transferase activity / positive regulation of protein targeting to mitochondrion / E2 ubiquitin-conjugating enzyme / cellular response to steroid hormone stimulus / immune system process / ubiquitin conjugating enzyme activity / protein K63-linked ubiquitination / regulation of lipid metabolic process / protein autoubiquitination / ubiquitin ligase complex / Synthesis of active ubiquitin: roles of E1 and E2 enzymes / lipid droplet / positive regulation of protein ubiquitination / Regulation of TNFR1 signaling / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / RING-type E3 ubiquitin transferase / protein modification process / Regulation of necroptotic cell death / protein polyubiquitination / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / Antigen processing: Ubiquitination & Proteasome degradation / presynapse / E3 ubiquitin ligases ubiquitinate target proteins / ubiquitin-dependent protein catabolic process / angiogenesis / cell population proliferation / transcription coactivator activity / protein ubiquitination / defense response to bacterium / ubiquitin protein ligase binding / regulation of DNA-templated transcription / nucleolus / enzyme binding / ATP hydrolysis activity / RNA binding / nucleoplasm / ATP binding / membrane / nucleus / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
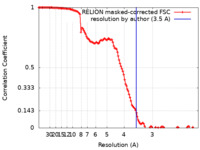
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||||||||
 Authors Authors | Ahel J / Clausen T | |||||||||||||||
| Funding support |  Austria, Austria,  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: E3 ubiquitin ligase RNF213 employs a non-canonical zinc finger active site and is allosterically regulated by ATP Authors: Ahel J / Fletcher AJ / Grabarczyk D / Roitinger E / Deszcz L / Lehner A / Virdee S / Clausen T | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12931.map.gz emd_12931.map.gz | 131.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12931-v30.xml emd-12931-v30.xml emd-12931.xml emd-12931.xml | 26.5 KB 26.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12931_fsc.xml emd_12931_fsc.xml | 12.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_12931.png emd_12931.png | 188.7 KB | ||
| Masks |  emd_12931_msk_1.map emd_12931_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Others |  emd_12931_half_map_1.map.gz emd_12931_half_map_1.map.gz emd_12931_half_map_2.map.gz emd_12931_half_map_2.map.gz | 131.7 MB 131.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12931 http://ftp.pdbj.org/pub/emdb/structures/EMD-12931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12931 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12931 | HTTPS FTP |
-Validation report
| Summary document |  emd_12931_validation.pdf.gz emd_12931_validation.pdf.gz | 765.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12931_full_validation.pdf.gz emd_12931_full_validation.pdf.gz | 765.4 KB | Display | |
| Data in XML |  emd_12931_validation.xml.gz emd_12931_validation.xml.gz | 19.8 KB | Display | |
| Data in CIF |  emd_12931_validation.cif.gz emd_12931_validation.cif.gz | 26.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12931 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12931 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12931 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12931 | HTTPS FTP |
-Related structure data
| Related structure data |  7oikMC  7oimC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-10711 (Title: Transmission electron micrographs of mouse RNF213 in a chemically-stabilized complex with human UBE2L3 EMPIAR-10711 (Title: Transmission electron micrographs of mouse RNF213 in a chemically-stabilized complex with human UBE2L3Data size: 2.1 TB Data #1: unaligned multi-frame micrographs [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12931.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12931.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | full reconstructed density | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.072 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_12931_msk_1.map emd_12931_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
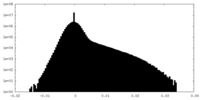
| Density Histograms |
-Half map: unfiltered half map 2 of the full reconstructed density
| File | emd_12931_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered half map 2 of the full reconstructed density | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: unfiltered half map 1 of the full reconstructed density
| File | emd_12931_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered half map 1 of the full reconstructed density | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : RNF213-ABP(UBE2L3-Ub)
| Entire | Name: RNF213-ABP(UBE2L3-Ub) |
|---|---|
| Components |
|
-Supramolecule #1: RNF213-ABP(UBE2L3-Ub)
| Supramolecule | Name: RNF213-ABP(UBE2L3-Ub) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: chemically stabilized complex between mouse RNF213 and a modified human UBE2L3 ABP (activity based probe), coupled to modified human ubiquitin |
|---|---|
| Molecular weight | Theoretical: 580 KDa |
-Supramolecule #2: RNF213-ABP(UBE2L3-Ub)
| Supramolecule | Name: RNF213-ABP(UBE2L3-Ub) / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 / Details: E3 ubiquitin-protein ligase RNF213 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
-Supramolecule #3: RNF213-ABP(UBE2L3-Ub)
| Supramolecule | Name: RNF213-ABP(UBE2L3-Ub) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 / Details: Ubiquitin-conjugating enzyme E2 L3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: E3 ubiquitin-protein ligase RNF213
| Macromolecule | Name: E3 ubiquitin-protein ligase RNF213 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RING-type E3 ubiquitin transferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 586.5455 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MECPQCGHVS SEKAPKFCSE CGQKLPSAAT VQGDLKNDNT LVVSSTPEGK TEQGAVLREE EVLLSSTDPG KELEKPEESD SNASWTTQM SKKEKRRRKR QGTISSSEAP SSGLWSLDMP PSPGSHNSAL PQNQAQQGGA ASQPGHPLDT ENMPMEDGFV H TEGSGSPL ...String: MECPQCGHVS SEKAPKFCSE CGQKLPSAAT VQGDLKNDNT LVVSSTPEGK TEQGAVLREE EVLLSSTDPG KELEKPEESD SNASWTTQM SKKEKRRRKR QGTISSSEAP SSGLWSLDMP PSPGSHNSAL PQNQAQQGGA ASQPGHPLDT ENMPMEDGFV H TEGSGSPL QGQAAERTDA QSNLAPSDLA EVKDLNTSKP SVDKGLPLDG GPALSAFKGH PKMTDASQKA PLPESKGETS GQ EKKVPPI DAAASPVKTA GKETGEDVRK PKPSPVSPVA SKHGDQEAEL KGKLATPVRK SNEGGNTQPE DQRKPGEGRN FAA AVKTQQ AAAPQQAAAP EPTSAFNPRD TVTVYFHAIV SRHFGFNPEE HKVYVRGGEG LGQKGWTDAC EMYCTQDLHD LGSL VEGKM DIPRQSLDKP IPYKYVIHRG GSSKDTVEYE FIYEQAQKKG EHVNRCLRVV STSLGNGDWH QYDDIICMRS TGFFQ QAKN RILDSTRKEL LKGKKQAAVV MLDRIFSVLQ PWSDINLQSF MTQFLQFYSV VREPMIHDGR ARKWTSLQYE EKEVWT NLW EHVKKQMAPF LEGKSGESLP ADCPVRSKLT LGLSILFMVE AAEFTVPKKD LDSLCYLLIP SAGSPEALHS DLSPVLR IR QRWRIYLTNL CLRCIDERCD RWLGILPLLH TCMQKSPPKK NSKSQPEDTW AGLEGISFSE FRDKAPTRSQ PLQFMQSK M ALLRVDEYLF RSWLSVVPLE SLSSYLENSI DYLSDVPVRV LDCLQGISYR LPGLRKISNQ NMKKDVENVF KMLMHLVDI YQHRIFGENL LQIYLTECLT LHETVCNITA NHQFFEIPAL SAELICKLLE LSPPGHTDEG LPEKSYEDLV TSTLQEALAT TRNWLRSLF KSRMLSISSA YVRLTYSEEM AVWRRLVEIG FPEKHGWKGS LLGDMEGRLK QEPPRLQISF FCSSQCRDGG L HDSVSRSF EKCVIEAVSS ACQSQTSVLE GLSCQDLQKF GTLLSAVITK SWPVHNGEPV FDVDEIFKYL LKWPDVRQLF EL CGTNEKI IDNITEEGRQ LMATAESVFQ KVAGELENGT IVVGQLELIL EHQSQFLDIW NLNRRRLPSQ EKACDVRSLL KRR RDDLLF LKQEKRYVES LLRQLGRVKH LVQVDFGNIE IIHSQDLSNK KLNEAVIKLP NSSSYKRETH YCLSPDIREM ASKL DSLKD SHIFQDFWQE TAESLNTLDK DPRELKVSLP EVLEYLYNPC YDNFYTLYEN LKSGKITFAE VDAIFKDFVD KYDEL KNDL KFMCTMNPQD QKGWISERVG QIKEYHTLHQ AVSSAKVILQ VRRALGVTGD FSVLNPLLNF ADSFEDFGNE KLDQIS PQF IKAKQLLQDI SEPRQRCLEE LARQTELVAW LHKALEDINE LKVFVDLASI SAGENDIDVD RVACFHDAVQ GYASLLY KM DERTNFSDFM NHLQELWRAL DNDQHLPDKL KDSARNLEWL KTVKESHGSV ELSSLSLATA INSRGVYVIE APKDGQKI S PDTVLRLLLP DGHGYPEALR TYSTEELKEL LNKLMLMSGK KDHNSNTEVE KFSEVFSNMQ RLVHVFIKLH CAGNMLFRT WTAKVYCCPD GGIFMNFGLE LLSQLTEKGD VIQLLGALCR QMEDFLDNWK TVVAQKRAEH FYLNFYTAEQ LVYLSSELRK PRPSEAALM MLSFIKGKCT VQDLVQATSA CESKADRYCL REVMKKLPQQ LLSEPSLMGK LQVIMMQSLV YMSAFLPHCL D LDALGRCL AHLATMGGTP VERPLPKGLQ AGQPNLILCG HSEVLPAALA IYMQAPRQPL PTFDEVLLCT PATTIEEVEL LL RRCLTSG SQGHKVYSLL FADQLSYEVG CQAEEFFQSL CTRAHREDYQ LVILCDAARE HCYIPSTFSQ YKVPLVPQAP LPN IQAYLQ SHYQVPKRLL SAATVFRDGL CVGIVTSERA GVGKSLYVNT LHTKLKAKLR DETVPLKIIR LTEPHLDENQ VLSA LLPFL KEKYQKMPVI FHIDISTSVQ TGIPIFLFKL LILQYLMDIN GKIWRRSPGH LYLVEIPQGL SVQPKRSSKL NARAP LFKF LDLFPKVTCR PPKEVIDMEL TPERSHTDPA MDPVEFCSEA FQRPYQYLKR FHQQQNLDTF QYEKGSVEGS PEECLQ HFL IYCGLINPSW SELRNFAWFL NCQLKDCEAS IFCKSAFTGD TLRGFKNFVV TFMILMARDF ATPTLHTSDQ SPGRQSV TI GEVVEEDLAP FSLRKRWESE PHPYVFFNGD HMTMTFIGFH LETNNNGYVD AINPSNGKVI KKDVMTKELF DGLRLQRV P FNIDFDNLPR YEKLERLCLA LGIEWPIDPD ETYELTTDNM LKILAIEMRF RCGIPVIIMG ETGCGKTRLI KFLSDLKRG SVEAETMKLV KVHGGTTPSM IYSKVKEAER TAFSNKAQHK LDTILFFDEA NTTEAVSCIK EILCDRTVDG EHLHEDSGLH IIAACNPYR KHSQEMILRL ESAGLGYRVS AEETADRLGS IPLRQLVYRV HALPPSLIPL VWDFGQLNDS AEKLYIQQIV Q RLVDSVSV NPSETCVIAD VLSASQMFMR KRENECGFVS LRDVERCVKV FRWFHDHSDM LLKELDKFLH ESSDSTHTFE RD PVLWSLV MAIGVCYHAS LEEKASYRTA IARCFPKPYN SSRAILDEVT HVQDLFLRGA PIRTNIARNL ALKENVFMMV ICI ELKIPL FLVGKPGSSK SLAKIIVADA MQGQAAFSEL FRCLKQVHLV SFQCSPHSTP QGIISTFKQC ARFQQGKDLG QYVS VVVLD EVGLAEDSPK MPLKTLHPLL EDGCIEDDPA PYKKVGFVGI SNWALDPAKM NRGIFVSRGS PNEKELIESA EGICS SDRL VQDKIRGYFA PFAKAYETVC QKQDKEFFGL RDYYSLIKMV FAKAKASKRG LSPQDITHAV LRNFSGKDNI QALSIF TAS LPEARYKEEV STVELIKQNI YPGPQASSRG LDGAESRYLL VLTRNYVALQ ILQQTFFEGQ QPEIIFGSSF PQDQEYT QI CRNINRVKIC METGKMVVLL NLQNLYESLY DALNQYYVYL GGQKYVDLGL GTHRVKCRVH TAFRLIVIEE KDVVYKQF P VPLINRLEKH YLDMNTVLQP WQKSIVQELQ QWAHEFADVK ADQFIARHKY SPADVFIGYH SDACASVVLQ AVERQGCRD LTEELYRKVS EEARSILLDC ATPDAVVRLS GSSLGSFTAK QLSQEYYYAQ QHNSFVDFLQ AHLRMTHHEC RAVFTEITTF SRLLTGNDC DVLASELRGL ASKPVVLSLQ QYDTEYSFLK DVRSWLTNPG KRKVLVIQAD FDDGTRSAQL VASAKYTAIN E INKTQGTK DFVFVYFVTK LSRMGSGTSY VGFHGGLWRS VHIDDLRRST IMASDVTKLQ NVTISQLFKP EDKPEQEEME IE TSQSKEL AEEQMEVEDS EEMKKASDPR SCDCSQFLDT TRLVQSCVQG AVGMLRDQNE SCARNMRRVT ILLDLLNEDN TRN ASFLRE SKMRLHVLLN KQEENQVRSL KEWVTREAAN QDALQEAGTF RHTLWKRVQD VVTPILASMI AHIDRDGNLE LLAQ PDSPA WVQDLWMFIY SDIKFLNISL VLNNTRSNSE MSFILVQSHM NLLKDAYNAV PFSWRIRDYL EELWVQAQYI TDTEG LSKK FVEIFQKTPL GVFLAQFPVA QQQKLLQSYL KDFLLLTMKV SSREELMFLQ MALWSCLREL QEASGTPDET YKFPLS LPW VHLAFQHFRT RLQNFSRILT IHPQVLSSLS QAAEKHSLAG CEMTLDAFAA MACAEMLKGD LLKPSPKAWL QLVKNLS TP LELVCSEGYL CDSGSMTRSV IQEVRALWNR IFSIALFVEH VLLGTESHIP ELSPLVTTYV SLLDKCLEED SNLKTCRP F VAVMTTLCDC KDKASKKFSR FGIQPCFICH GDAQDPVCLP CDHVYCLRCI QTWLIPGQMM CPYCLTDLPD KFSPTVSQD HRKAIEKHAQ FRHMCNSFFV DLVSTMCFKD NTPPEKSVID TLLSLLFVQK ELLRDASQKH REHTKSLSPF DDVVDQTPVI RSVLLKLLL KYSFHEVKDY IQNYLTQLEK KAFLTEDKTE LYLLFISCLE DSVHQKTSAG CRNLEQVLRE EGHFLRTYSP G LQGQEPVR IASVEYLQEV ARVRLCLDLA ADFLSELQEG SELAEDKRRF LKHVEEFCTR VNNDWHRVYL VRKLSSQRGM EF VQSFSKQ GHPCQWVFPR KVIAQQKDHV SLMDRYLVHG NEYKAVRDAT AKAVLECKTL DIGNALMACR SPKPQQTAYL LLA LYTEVA ALYRSPNGSL HPEAKQLEAV NKFIKESKIL SDPNIRCFAR SLVDNTLPLL KIRSANSILK GTVTEMAVHV ATIL LCGHN QILKPLRNLA FYPVNMANAF LPTMPEDLLV HARTWRGLEN VTWYTCPRGH PCSVGECGRP MQESTCLDCG LPVGG LNHT PHEGFSAIRN NEDRTQTGHV LGSPQSSGVA EVSDRGQSPV VFILTRLLTH LAMLVGATHN PQALTVIIKP WVQDPQ GFL QQHIQRDLEQ LTKMLGRSAD ETIHVVHLIL SSLLRVQSHG VLNFNAELST KGCRNNWEKH FETLLLRELK HLDKNLP AI NALISQDERI SSNPVTKIIY GDPATFLPHL PQKSIIHCSK IWSCRRKITV EYLQHIVEQK NGKETVPVLW HFLQKEAE L RLVKFLPEIL ALQRDLVKQF QNVSRVEYSS IRGFIHSHSS DGLRKLLHDR ITIFLSTWNA LRRSLETNGE IKLPKDYCC SDLDLDAEFE VILPRRQGLG LCGTALVSYL ISLHNNMVYT VQKFSNEDNS YSVDISEVAD LHVISYEVER DLNPLILSNC QYQVQQGGE TSQEFDLEKI QRQISSRFLQ GKPRLTLKGI PTLVYRRDWN YEHLFMDIKN KMAQSSLPNL AISTISGQLQ S YSDACEAL SIIEITLGFL STAGGDPGMD LNVYIEEVLR MCDQTAQVLK AFSRCQLRHI IALWQFLSAH KSEQRLRLNK EL FREIDVQ YKEELSTQHQ RLLGTFLNEA GLDAFLLELH EMIVLKLKGP RAANSFNPNW SLKDTLVSYM ETKDSDILSE VES QFPEEI LMSSCISVWK IAATRKWDRQ SRGGGHHHHH HHHHH |
-Macromolecule #2: Ubiquitin-conjugating enzyme E2 L3
| Macromolecule | Name: Ubiquitin-conjugating enzyme E2 L3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: E2 ubiquitin-conjugating enzyme |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 19.197908 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GWSHPQFEKP GSMAASRRLM KELEEIRKSG MKNFRNIQVD EANLLTWQGL IVPDNPPYDK GAFRIEINFP AEYPFKPPKI TFKTKIYHP NIDEKGQVCL PVISAENWKP ATKTDQVIQS LIALVNDPQP EHPLRADLAE EYSKDRKKFS KNAEEFTKKY G EKRPVD |
-Macromolecule #3: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 1 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil R2/2 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 4591 / Average exposure time: 3.92 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -2.5 µm / Nominal defocus min: -0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X