+ Open data
Open data
- Basic information
Basic information
| Entry |  Database: PDB chemical components / ID: DM5 Database: PDB chemical components / ID: DM5 |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAD / Three letter code: DM5 / Model coordinates PDB-ID: 1D67 | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB / BindingDB /  Brenda / Brenda /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  HMDB / HMDB /  PubChem / PubChem /  SureChEMBL SureChEMBL   / /  Wikipedia search / Wikipedia search /  Google search Google search |
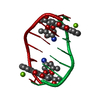
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | (| OpenEye OEToolkits 1.5.0 | ( | |
|---|
-PDB entries
Showing all 5 items

PDB-198d: 
A TRIGONAL FORM OF THE IDARUBICIN-D(CGATCG) COMPLEX: CRYSTAL AND MOLECULAR STRUCTURE AT 2.0 ANGSTROMS RESOLUTION
PDB-1d38: 
INFLUENCE OF AGLYCONE MODIFICATIONS ON THE BINDING OF ANTHRACYCLINE DRUGS TO DNA: THE MOLECULAR STRUCTURE OF IDARUBICIN AND 4-O-DEMETHYL-11-DEOXYDOXORUBICIN COMPLEXED TO D(CGATCG)

PDB-1d67: 
THE MOLECULAR STRUCTURE OF AN IDARUBICIN-D(TGATCA) COMPLEX AT HIGH RESOLUTION

PDB-3arq: 
Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with IDARUBICIN

PDB-4lb2: 
X-ray study of human serum albumin complexed with idarubicin
 Movie
Movie Controller
Controller



