+ Open data
Open data
- Basic information
Basic information
| Entry | 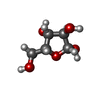 Database: PDB chemical components / ID: XYZ Database: PDB chemical components / ID: XYZ |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Others | Type: D-saccharide, beta linking / PDB classification: ATOMS / Three letter code: XYZ / Model coordinates PDB-ID: 1W0Q | ||||||||||||||||||
| History |
| ||||||||||||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda / Brenda /  ChEBI / ChEBI /  PubChem / PubChem /  SureChEMBL / SureChEMBL /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | ( | |
|---|
-CONDENSED IUPAC CARBOHYDRATE SYMBOL
| GMML 1.0 |
|---|
-COMMON NAME
| GMML 1.0 |
|---|
-IUPAC CARBOHYDRATE SYMBOL
| PDB-CARE 1.0 |
|---|
-SNFG CARBOHYDRATE SYMBOL
| GMML 1.0 |
|---|
-PDB entries
Showing all 5 items

PDB-5xis: 
Crystal structure of RNF168 UDM1 in complex with Lys63-linked diubiquitin, form I

PDB-7ey1: 
Bifunctional xylosidase/glucosidase LXYL with intermediate substrate xylose

PDB-7ey2: 
Bifunctional xylosidase/glucosidase LXYL D300N mutant with intermediate substrate xylose

PDB-7rte: 
X-ray structure of wild type RBPJ-L3MBTL3-DNA complex

PDB-7yo6: 
Bifunctional xylosidase/glucosidase LXYL with intermediate substrate xylose for 1.5min
 Movie
Movie Controller
Controller




