+ Open data
Open data
- Basic information
Basic information
| Entry | 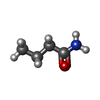 Database: PDB chemical components / ID: BMD Database: PDB chemical components / ID: BMD |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: BMD / Model coordinates PDB-ID: 1QO0 | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda Brenda     / /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  HMDB / HMDB /  LipidMaps / LipidMaps /  PubChem / PubChem /  Rhea / Rhea /  SureChEMBL SureChEMBL  / /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 4 items

PDB-1qnl: 
AMIDE RECEPTOR/NEGATIVE REGULATOR OF THE AMIDASE OPERON OF PSEUDOMONAS AERUGINOSA (AMIC) COMPLEXED WITH BUTYRAMIDE

PDB-1qo0: 
Amide receptor of the amidase operon of Pseudomonas aeruginosa (AmiC) complexed with the negative regulator AmiR.

PDB-3pii: 
Crystal structure of Mutant of ht- Alcohol Dehydrogenase with substrate analogue butyramide

PDB-4izs: 
The C145A mutant of the amidase from Nesterenkonia sp. AN1 in complex with butyramide
 Movie
Movie Controller
Controller



