+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9205 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
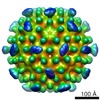
| Title | Red Clover Necrotic Mosaic Virus | |||||||||
 Map data Map data | Red clover necrotic mosaic virus | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RCNMV / virus | |||||||||
| Function / homology | Plant viruses icosahedral capsid proteins 'S' region signature. / Icosahedral viral capsid protein, S domain / Viral coat protein (S domain) / T=3 icosahedral viral capsid / Viral coat protein subunit / structural molecule activity / RNA binding / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Red clover necrotic mosaic virus Red clover necrotic mosaic virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Sherman MB / Smith TJ | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2020 Journal: J Virol / Year: 2020Title: Near-Atomic-Resolution Cryo-Electron Microscopy Structures of Cucumber Leaf Spot Virus and Red Clover Necrotic Mosaic Virus: Evolutionary Divergence at the Icosahedral Three-Fold Axes. Authors: Michael B Sherman / Richard Guenther / Ron Reade / D'Ann Rochon / Tim Sit / Thomas J Smith /   Abstract: Members of the family have highly similar structures, and yet there are important differences among them in host, transmission, and capsid stabilities. Viruses in the family have single-stranded ...Members of the family have highly similar structures, and yet there are important differences among them in host, transmission, and capsid stabilities. Viruses in the family have single-stranded RNA (ssRNA) genomes with T=3 icosahedral protein shells with a maximum diameter of ∼340 Å. Each capsid protein is comprised of three domains: R (RNA binding), S (shell), and P (protruding). Between the R domain and S domain is the "arm" region that studies have shown to play a critical role in assembly. To better understand how the details of structural differences and similarities influence the viral life cycles, the structures of cucumber leaf spot virus (CLSV; genus ) and red clover necrotic mosaic virus (RCNMV; genus ) were determined to resolutions of 3.2 Å and 2.9 Å, respectively, with cryo-electron microscopy and image reconstruction methods. While the shell domains had homologous structures, the stabilizing interactions at the icosahedral 3-fold axes and the R domains differed greatly. The heterogeneity in the R domains among the members of the family is likely correlated with differences in the sizes and characteristics of the corresponding genomes. We propose that the changes in the R domain/RNA interactions evolved different arm domain interactions at the β-annuli. For example, RCNMV has the largest genome and it appears to have created the necessary space in the capsid by evolving the shortest R domain. The resulting loss in RNA/R domain interactions may have been compensated for by increased intersubunit β-strand interactions at the icosahedral 3-fold axes. Therefore, the R and arm domains may have coevolved to package different genomes within the conserved and rigid shell. Members of the family have nearly identical shells, and yet they package genomes that range from 4.6 kb (monopartite) to 5.3 kb (bipartite) in size. To understand how this genome flexibility occurs within a rigidly conserved shell, we determined the high-resolution cryo-electron microscopy (cryo-EM) structures of cucumber leaf spot virus and red clover necrotic mosaic virus. In response to genomic size differences, it appears that the ssRNA binding (R) domain of the capsid diverged evolutionarily in order to recognize the different genomes. The next region, the "arm," seems to have also coevolved with the R domain to allow particle assembly via interactions at the icosahedral 3-fold axes. In addition, there are differences at the icosahedral 3-fold axes with regard to metal binding that are likely important for transmission and the viral life cycle. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9205.map.gz emd_9205.map.gz | 476.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9205-v30.xml emd-9205-v30.xml emd-9205.xml emd-9205.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9205.png emd_9205.png | 261.4 KB | ||
| Filedesc metadata |  emd-9205.cif.gz emd-9205.cif.gz | 5.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9205 http://ftp.pdbj.org/pub/emdb/structures/EMD-9205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9205 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9205 | HTTPS FTP |
-Validation report
| Summary document |  emd_9205_validation.pdf.gz emd_9205_validation.pdf.gz | 698.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9205_full_validation.pdf.gz emd_9205_full_validation.pdf.gz | 698.2 KB | Display | |
| Data in XML |  emd_9205_validation.xml.gz emd_9205_validation.xml.gz | 8.7 KB | Display | |
| Data in CIF |  emd_9205_validation.cif.gz emd_9205_validation.cif.gz | 10 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9205 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9205 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9205 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9205 | HTTPS FTP |
-Related structure data
| Related structure data |  6mrmMC  9204C  6mrlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9205.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9205.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Red clover necrotic mosaic virus | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0556 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Red clover necrotic mosaic virus
| Entire | Name:  Red clover necrotic mosaic virus Red clover necrotic mosaic virus |
|---|---|
| Components |
|
-Supramolecule #1: Red clover necrotic mosaic virus
| Supramolecule | Name: Red clover necrotic mosaic virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / NCBI-ID: 12267 / Sci species name: Red clover necrotic mosaic virus / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Red clover necrotic mosaic virus Red clover necrotic mosaic virus |
| Molecular weight | Theoretical: 36.617473 KDa |
| Sequence | String: MSSKAPKKSK QRSQPRNRTP NTSVKTVAIP FAKTQIIKTV NPPPKPARGI LHTQLVMSVV GSVQMRTNNG KSNQRFRLNP SNPALFPTL AYEAANYDMY RLKKLTLRYV PLVTVQNSGR VAMIWDPDSQ DSAPQSRQEI SAYSRSVSTA VYEKCSLTIP A DNQWRFVA ...String: MSSKAPKKSK QRSQPRNRTP NTSVKTVAIP FAKTQIIKTV NPPPKPARGI LHTQLVMSVV GSVQMRTNNG KSNQRFRLNP SNPALFPTL AYEAANYDMY RLKKLTLRYV PLVTVQNSGR VAMIWDPDSQ DSAPQSRQEI SAYSRSVSTA VYEKCSLTIP A DNQWRFVA DNTTVDRKLV DFGQLLFVTH SGSDGIETGD IFLDCEVEFK GPQPTASIVQ KTVIDLGGTL TSFEGPSYLM PP DAFITSS SFGLFVDVAG TYLLTLVVTC STTGSVTVGG NSTLVGDGRA AYGSSNYIAS IVFTSSGVLS TTPSVQFSGS SGV SRVQMN ICRCKQGNTF ILG UniProtKB: Capsid protein |
-Macromolecule #2: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 2 / Number of copies: 3 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: DIRECT ELECTRON DE-64 (8k x 8k) / Average electron dose: 54.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1236 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















