+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6789 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Large subunit of Mycobacterium smegmatis | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mycobacterium smegmatis ribosome / rRNA / rprotein / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology informationlarge ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding ...large ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.08 Å | |||||||||
 Authors Authors | Li Z / Ge X | |||||||||
 Citation Citation |  Journal: Protein Cell / Year: 2018 Journal: Protein Cell / Year: 2018Title: Cryo-EM structure of Mycobacterium smegmatis ribosome reveals two unidentified ribosomal proteins close to the functional centers. Authors: Zhifei Li / Xueliang Ge / Yixiao Zhang / Lvqin Zheng / Suparna Sanyal / Ning Gao /   | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6789.map.gz emd_6789.map.gz | 9.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6789-v30.xml emd-6789-v30.xml emd-6789.xml emd-6789.xml | 43.2 KB 43.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6789_fsc.xml emd_6789_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_6789.png emd_6789.png | 95 KB | ||
| Filedesc metadata |  emd-6789.cif.gz emd-6789.cif.gz | 9.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6789 http://ftp.pdbj.org/pub/emdb/structures/EMD-6789 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6789 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6789 | HTTPS FTP |
-Related structure data
| Related structure data |  5xymMC  6790C  5xyuC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6789.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6789.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
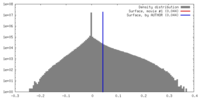
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Large subunit of Mycobacterium smegmatis ribosome
+Supramolecule #1: Large subunit of Mycobacterium smegmatis ribosome
+Macromolecule #1: 50S ribosomal protein L32
+Macromolecule #2: 50S ribosomal protein L33 2
+Macromolecule #3: 50S ribosomal protein L34
+Macromolecule #4: 50S ribosomal protein L35
+Macromolecule #5: 50S ribosomal protein L36
+Macromolecule #8: 50S ribosomal protein L2
+Macromolecule #9: 50S ribosomal protein L3
+Macromolecule #10: 50S ribosomal protein L4
+Macromolecule #11: 50S ribosomal protein L5
+Macromolecule #12: 50S ribosomal protein L6
+Macromolecule #13: 50S ribosomal protein L9
+Macromolecule #14: 50S ribosomal protein L13
+Macromolecule #15: 50S ribosomal protein L14
+Macromolecule #16: 50S ribosomal protein L15
+Macromolecule #17: 50S ribosomal protein L16
+Macromolecule #18: 50S ribosomal protein L17
+Macromolecule #19: 50S ribosomal protein L18
+Macromolecule #20: 50S ribosomal protein L19
+Macromolecule #21: 50S ribosomal protein L20
+Macromolecule #22: 50S ribosomal protein L21
+Macromolecule #23: 50S ribosomal protein L22
+Macromolecule #24: 50S ribosomal protein L23
+Macromolecule #25: 50S ribosomal protein L24
+Macromolecule #26: 50S ribosomal protein L25
+Macromolecule #27: 50S ribosomal protein L27
+Macromolecule #28: 50S ribosomal protein L28
+Macromolecule #29: 50S ribosomal protein L29
+Macromolecule #30: 50S ribosomal protein L30
+Macromolecule #31: Uncharacterized protein bL37
+Macromolecule #6: 23S RNA
+Macromolecule #7: 5S RNA
+Macromolecule #32: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 2.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)