[English] 日本語
 Yorodumi
Yorodumi- EMDB-6387: A new view of essential vertex proteins on herpesvirus capsids ob... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6387 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A new view of essential vertex proteins on herpesvirus capsids observed inside intact virions | |||||||||
 Map data Map data | Reconstruction of pseudorabies virus capsid imaged inside virions | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Pseudorabies virus / PRV / capsid / virion / CVSC / pUL17 / pUL25 / pUL36 | |||||||||
| Biological species |   Suid herpesvirus 1 Suid herpesvirus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.2 Å | |||||||||
 Authors Authors | Huet A / Makhov AM / Huffman JB / Vos M / Homa FL / Conway JF | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2016 Journal: Nat Struct Mol Biol / Year: 2016Title: Extensive subunit contacts underpin herpesvirus capsid stability and interior-to-exterior allostery. Authors: Alexis Huet / Alexander M Makhov / Jamie B Huffman / Matthijn Vos / Fred L Homa / James F Conway /   Abstract: The herpesvirus capsid is a complex protein assembly that includes hundreds of copies of four major subunits and lesser numbers of several minor proteins, all of which are essential for infectivity. ...The herpesvirus capsid is a complex protein assembly that includes hundreds of copies of four major subunits and lesser numbers of several minor proteins, all of which are essential for infectivity. Cryo-electron microscopy is uniquely suited for studying interactions that govern the assembly and function of such large functional complexes. Here we report two high-quality capsid structures, from human herpes simplex virus type 1 (HSV-1) and the animal pseudorabies virus (PRV), imaged inside intact virions at ~7-Å resolution. From these, we developed a complete model of subunit and domain organization and identified extensive networks of subunit contacts that underpin capsid stability and form a pathway that may signal the completion of DNA packaging from the capsid interior to outer surface, thereby initiating nuclear egress. Differences in the folding and orientation of subunit domains between herpesvirus capsids suggest that common elements have been modified for specific functions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6387.map.gz emd_6387.map.gz | 389.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6387-v30.xml emd-6387-v30.xml emd-6387.xml emd-6387.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6387.tif emd_6387.tif emd_6387_1.tif emd_6387_1.tif | 5.2 MB 2 MB | ||
| Others |  EMD-6387_1.tif EMD-6387_1.tif | 2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6387 http://ftp.pdbj.org/pub/emdb/structures/EMD-6387 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6387 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6387 | HTTPS FTP |
-Validation report
| Summary document |  emd_6387_validation.pdf.gz emd_6387_validation.pdf.gz | 77.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6387_full_validation.pdf.gz emd_6387_full_validation.pdf.gz | 76.9 KB | Display | |
| Data in XML |  emd_6387_validation.xml.gz emd_6387_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6387 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6387 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6387 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6387 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6387.map.gz / Format: CCP4 / Size: 782.7 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_6387.map.gz / Format: CCP4 / Size: 782.7 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of pseudorabies virus capsid imaged inside virions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
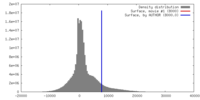
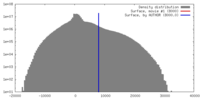
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Others
| Image |
|---|
- Sample components
Sample components
-Entire : Capsid of pseudorabies virus imaged in intact virions
| Entire | Name: Capsid of pseudorabies virus imaged in intact virions |
|---|---|
| Components |
|
-Supramolecule #1000: Capsid of pseudorabies virus imaged in intact virions
| Supramolecule | Name: Capsid of pseudorabies virus imaged in intact virions / type: sample / ID: 1000 / Oligomeric state: Icosahedral T=16 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 200 MDa / Theoretical: 200 MDa |
-Supramolecule #1: Suid herpesvirus 1
| Supramolecule | Name: Suid herpesvirus 1 / type: virus / ID: 1 / Details: Native Becker / NCBI-ID: 10345 / Sci species name: Suid herpesvirus 1 / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Host system | Organism:  Chlorocebus aethiops (grivet) / Recombinant cell: green monkey kidney (Vero) cells Chlorocebus aethiops (grivet) / Recombinant cell: green monkey kidney (Vero) cells |
| Molecular weight | Experimental: 200 MDa / Theoretical: 200 MDa |
| Virus shell | Shell ID: 1 / Diameter: 1250 Å / T number (triangulation number): 16 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Grid | Details: Quantifoil R2/1 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 120 K / Instrument: FEI VITROBOT MARK IV / Method: 6 second blot |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 75,000x magnification. |
| Details | Nanoprobe |
| Date | Apr 14, 2013 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON II (4k x 4k) / Digitization - Sampling interval: 14 µm / Number real images: 32956 / Average electron dose: 30 e/Å2 / Details: 2 second exposures; no use of movie mode |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 81000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder: Nitrogen cooled / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Manual selection, processing with AUTO3DEM |
|---|---|
| CTF correction | Details: By micrograph |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.2 Å / Resolution method: OTHER / Software - Name: AUTO3DEM, CTFFIND3 Details: Final map calculated from 25637 images of capsids inside virions. Resolution was verified by comparing cryoEM data with band-limited renditions of the HSV VP5 upper domain fragment, PDB 1NO7. Number images used: 13242 |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | Manual docking |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)