Entry Database : PDB / ID : 5kq5Title AMPK bound to allosteric activator (5'-AMP-activated protein kinase subunit ...) x 2 5'-AMP-activated protein kinase catalytic subunit alpha-1 Keywords / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Rattus norvegicus (Norway rat)Method / / / Resolution : 3.41 Å Authors Calabrese, M.F. / Kurumbail, R.G. Journal : J.Med.Chem. / Year : 2016Title: Discovery and Preclinical Characterization of 6-Chloro-5-[4-(1-hydroxycyclobutyl)phenyl]-1H-indole-3-carboxylic Acid (PF-06409577), a Direct Activator of Adenosine Monophosphate-activated ... Title : Discovery and Preclinical Characterization of 6-Chloro-5-[4-(1-hydroxycyclobutyl)phenyl]-1H-indole-3-carboxylic Acid (PF-06409577), a Direct Activator of Adenosine Monophosphate-activated Protein Kinase (AMPK), for the Potential Treatment of Diabetic Nephropathy.Authors: Cameron, K.O. / Kung, D.W. / Kalgutkar, A.S. / Kurumbail, R.G. / Miller, R. / Salatto, C.T. / Ward, J. / Withka, J.M. / Bhattacharya, S.K. / Boehm, M. / Borzilleri, K.A. / Brown, J.A. / ... Authors : Cameron, K.O. / Kung, D.W. / Kalgutkar, A.S. / Kurumbail, R.G. / Miller, R. / Salatto, C.T. / Ward, J. / Withka, J.M. / Bhattacharya, S.K. / Boehm, M. / Borzilleri, K.A. / Brown, J.A. / Calabrese, M. / Caspers, N.L. / Cokorinos, E. / Conn, E.L. / Dowling, M.S. / Edmonds, D.J. / Eng, H. / Fernando, D.P. / Frisbie, R. / Hepworth, D. / Landro, J. / Mao, Y. / Rajamohan, F. / Reyes, A.R. / Rose, C.R. / Ryder, T. / Shavnya, A. / Smith, A.C. / Tu, M. / Wolford, A.C. / Xiao, J. History Deposition Jul 5, 2016 Deposition site / Processing site Revision 1.0 Aug 17, 2016 Provider / Type Revision 1.1 Sep 21, 2016 Group Revision 1.2 Nov 1, 2017 Group / Database references / Derived calculationsCategory / pdbx_struct_assembly_auth_evidence / pdbx_struct_oper_listItem / _pdbx_struct_oper_list.symmetry_operationRevision 1.3 Oct 4, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accessionRevision 1.4 Oct 16, 2024 Group / Category / pdbx_modification_feature
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 3.41 Å
MOLECULAR REPLACEMENT / Resolution: 3.41 Å  Authors
Authors Citation
Citation Journal: J.Med.Chem. / Year: 2016
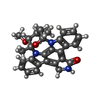
Journal: J.Med.Chem. / Year: 2016 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5kq5.cif.gz
5kq5.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5kq5.ent.gz
pdb5kq5.ent.gz PDB format
PDB format 5kq5.json.gz
5kq5.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/kq/5kq5
https://data.pdbj.org/pub/pdb/validation_reports/kq/5kq5 ftp://data.pdbj.org/pub/pdb/validation_reports/kq/5kq5
ftp://data.pdbj.org/pub/pdb/validation_reports/kq/5kq5
 Links
Links Assembly
Assembly
 Components
Components















 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 23-ID-D / Wavelength: 1 Å
/ Beamline: 23-ID-D / Wavelength: 1 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj











