| Entry | Database: PDB / ID: 4zaa
|
|---|
| Title | Structure of A. niger Fdc1 in complex with 4-vinyl guaiacol |
|---|
 Components Components | fdc1 |
|---|
 Keywords Keywords | LYASE / (de)carboxylase / prenylated flavin binding / UbiD-type enzyme |
|---|
| Function / homology |  Function and homology information Function and homology information
styrene metabolic process / aromatic amino acid catabolic process / phenacrylate decarboxylase / ferulate metabolic process / cinnamic acid catabolic process / carboxy-lyase activity / manganese ion binding / identical protein binding / cytoplasmSimilarity search - Function UbiD-like decarboxylase/ferulic acid decarboxylase 1 / : / : / : / 3-octaprenyl-4-hydroxybenzoate carboxy-lyase N-terminal domain / 3-octaprenyl-4-hydroxybenzoate carboxy-lyase C-terminal domain / UbiD decarboxylyase family / 3-octaprenyl-4-hydroxybenzoate carboxy-lyase Rift-related domainSimilarity search - Domain/homology |
|---|
| Biological species |   Aspergillus niger (mold) Aspergillus niger (mold) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.242 Å MOLECULAR REPLACEMENT / Resolution: 1.242 Å |
|---|
 Authors Authors | Payne, K.A.P. / Leys, D. |
|---|
 Citation Citation |  Journal: Nature / Year: 2015 Journal: Nature / Year: 2015
Title: New cofactor supports alpha , beta-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition.
Authors: Payne, K.A. / White, M.D. / Fisher, K. / Khara, B. / Bailey, S.S. / Parker, D. / Rattray, N.J. / Trivedi, D.K. / Goodacre, R. / Beveridge, R. / Barran, P. / Rigby, S.E. / Scrutton, N.S. / Hay, S. / Leys, D. |
|---|
| History | | Deposition | Apr 13, 2015 | Deposition site: RCSB / Processing site: PDBE |
|---|
| Revision 1.0 | Jun 17, 2015 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Jul 1, 2015 | Group: Database references |
|---|
| Revision 1.2 | May 8, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Structure summary
Category: chem_comp / chem_comp_atom ...chem_comp / chem_comp_atom / chem_comp_bond / database_2 / entity / pdbx_entity_nonpoly / pdbx_struct_conn_angle / struct_conn
Item: _chem_comp.name / _database_2.pdbx_DOI ..._chem_comp.name / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _entity.pdbx_description / _pdbx_entity_nonpoly.name / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.242 Å
MOLECULAR REPLACEMENT / Resolution: 1.242 Å  Authors
Authors Citation
Citation Journal: Nature / Year: 2015
Journal: Nature / Year: 2015 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4zaa.cif.gz
4zaa.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4zaa.ent.gz
pdb4zaa.ent.gz PDB format
PDB format 4zaa.json.gz
4zaa.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/za/4zaa
https://data.pdbj.org/pub/pdb/validation_reports/za/4zaa ftp://data.pdbj.org/pub/pdb/validation_reports/za/4zaa
ftp://data.pdbj.org/pub/pdb/validation_reports/za/4zaa







 Links
Links Assembly
Assembly

 Components
Components


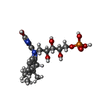


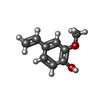







 X-RAY DIFFRACTION
X-RAY DIFFRACTION Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Diamond
Diamond  / Beamline: I24 / Wavelength: 0.98 Å
/ Beamline: I24 / Wavelength: 0.98 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.242→94.19 Å / Cor.coef. Fo:Fc: 0.975 / Cor.coef. Fo:Fc free: 0.971 / SU B: 1.327 / SU ML: 0.026 / Cross valid method: THROUGHOUT / ESU R: 0.038 / ESU R Free: 0.037 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.242→94.19 Å / Cor.coef. Fo:Fc: 0.975 / Cor.coef. Fo:Fc free: 0.971 / SU B: 1.327 / SU ML: 0.026 / Cross valid method: THROUGHOUT / ESU R: 0.038 / ESU R Free: 0.037 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller
















 PDBj
PDBj





