[English] 日本語
 Yorodumi
Yorodumi- EMDB-4997: Cryo-EM density of murine Solute Carrier 26 family member A9 (Slc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4997 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
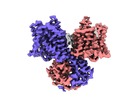
| Title | Cryo-EM density of murine Solute Carrier 26 family member A9 (Slc26a9) anion transporter in the inward-facing state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SLC26 family / anion transporter / membrane protein structure / transport mechanism / cryo-EM / single particle / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationInorganic anion exchange by SLC26 transporters / oxalate transmembrane transporter activity / sulfate transmembrane transporter activity / sulfate transmembrane transport / regulation of pH / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity / bicarbonate transmembrane transporter activity / bicarbonate transport / monoatomic anion transport ...Inorganic anion exchange by SLC26 transporters / oxalate transmembrane transporter activity / sulfate transmembrane transporter activity / sulfate transmembrane transport / regulation of pH / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity / bicarbonate transmembrane transporter activity / bicarbonate transport / monoatomic anion transport / chloride transport / chloride channel activity / chloride transmembrane transport / ATPase binding / endosome membrane / apical plasma membrane / Golgi membrane / positive regulation of gene expression / endoplasmic reticulum membrane / cell surface / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
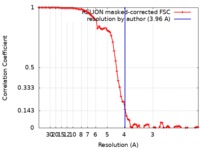
| Method | single particle reconstruction / cryo EM / Resolution: 3.96 Å | |||||||||
 Authors Authors | Sawicka M / Walter JD | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Cryo-EM structures and functional characterization of murine Slc26a9 reveal mechanism of uncoupled chloride transport. Authors: Justin D Walter / Marta Sawicka / Raimund Dutzler /  Abstract: The epithelial anion transporter SLC26A9 contributes to airway surface hydration and gastric acid production. Colocalizing with CFTR, SLC26A9 has been proposed as a target for the treatment of cystic ...The epithelial anion transporter SLC26A9 contributes to airway surface hydration and gastric acid production. Colocalizing with CFTR, SLC26A9 has been proposed as a target for the treatment of cystic fibrosis. To provide molecular details of its transport mechanism, we present cryo-EM structures and a functional characterization of murine Slc26a9. These structures define the general architecture of eukaryotic SLC26 family members and reveal an unusual mode of oligomerization which relies predominantly on the cytosolic STAS domain. Our data illustrates conformational transitions of Slc26a9, supporting a rapid alternate-access mechanism which mediates uncoupled chloride transport with negligible bicarbonate or sulfate permeability. The characterization of structure-guided mutants illuminates the properties of the ion transport path, including a selective anion binding site located in the center of a mobile module within the transmembrane domain. This study thus provides a structural foundation for the understanding of the entire SLC26 family and potentially facilitates their therapeutic exploitation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4997.map.gz emd_4997.map.gz | 3.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4997-v30.xml emd-4997-v30.xml emd-4997.xml emd-4997.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4997_fsc.xml emd_4997_fsc.xml | 8.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_4997.png emd_4997.png | 138.4 KB | ||
| Masks |  emd_4997_msk_1.map emd_4997_msk_1.map | 47.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-4997.cif.gz emd-4997.cif.gz | 6.7 KB | ||
| Others |  emd_4997_half_map_1.map.gz emd_4997_half_map_1.map.gz emd_4997_half_map_2.map.gz emd_4997_half_map_2.map.gz | 35.5 MB 35.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4997 http://ftp.pdbj.org/pub/emdb/structures/EMD-4997 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4997 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4997 | HTTPS FTP |
-Related structure data
| Related structure data |  6rtcMC  4998C  6rtfC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4997.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4997.map.gz / Format: CCP4 / Size: 47.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.075 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
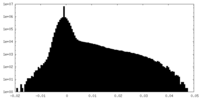
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4997_msk_1.map emd_4997_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
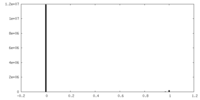
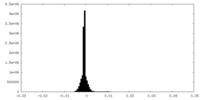
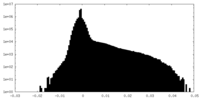
| Density Histograms |
-Half map: #2
| File | emd_4997_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
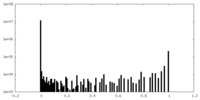
| Density Histograms |
-Half map: #1
| File | emd_4997_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
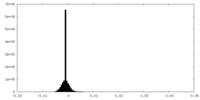
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimer of Solute Carrier 26 family member A9 (Slc26a9)
| Entire | Name: Homodimer of Solute Carrier 26 family member A9 (Slc26a9) |
|---|---|
| Components |
|
-Supramolecule #1: Homodimer of Solute Carrier 26 family member A9 (Slc26a9)
| Supramolecule | Name: Homodimer of Solute Carrier 26 family member A9 (Slc26a9) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Solute carrier family 26 member 9,Solute carrier family 26 member 9
| Macromolecule | Name: Solute carrier family 26 member 9,Solute carrier family 26 member 9 type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 70.597641 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MNQPRPRYVV DRAAYSLSLF DDEFEKKDRA YPVGEKLRNT FRCSSAKFKA FVFGLLPVLS WLPKYKIKDY IIPDLLGGLS GGCIQVPQG MAFALLANLP AVNGLYSSFF PLLTYFFLGG IHQMVPGTFA VISILVGNIC LQLAPESKFQ IFNNVTNETY V DTAAMEAE ...String: MNQPRPRYVV DRAAYSLSLF DDEFEKKDRA YPVGEKLRNT FRCSSAKFKA FVFGLLPVLS WLPKYKIKDY IIPDLLGGLS GGCIQVPQG MAFALLANLP AVNGLYSSFF PLLTYFFLGG IHQMVPGTFA VISILVGNIC LQLAPESKFQ IFNNVTNETY V DTAAMEAE RLHVSATLAC LTAVIQMALG FMQFGFVAIY LSESFIRGFM TAAGLQILIS VLKYIFGLTI PSYTGPGSIV FT FIDICKN LPHTNIASLI FALVSGVFLV LVKELNARYM HKIHFPIPTE MIVVVVATAI SGSCKMPKKY HMQIVGEIRQ GFP TPVAPM VSQWKDMVGT AFSLAIVGYV INLAMGRTLA SKHGYDVDSN QEMIALGCSN FFGSFFKIHV ICCALSVTLA VDGA GGKSQ VASLCVSLVV MITMLVLGSY LYPLPKAVLG ALIAVNLKNS LKQLTDPYYL WRKSKLDCCV WVVSFLSSFF LSLPY GVAV GVAFSILVVI FQTQFRNGST LAQVMDTDIY VNPKTYNRAQ EIAGVKIVTY CSPLYFANSE IFRQKVIAKT GMDGST FHT LILDMSGVSF VDLMGIKALA KLSSTYEKIG VQIFLVNIHA QVYNDISHGG VFEDGCVQRS HVFPSIHDAV LFAQANA RE A UniProtKB: Solute carrier family 26, member 9, Solute carrier family 26, member 9 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 2838 / Average exposure time: 0.2 sec. / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 46511 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 46511 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)