+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4657 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the human ClC-1 chloride channel, low pH | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | chloride channel / CLC1 / CLCN1 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationvoltage-gated chloride channel activity / neuronal action potential propagation / chloride transport / chloride channel complex / T-tubule / chloride transmembrane transport / muscle contraction / Stimuli-sensing channels / protein homodimerization activity / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
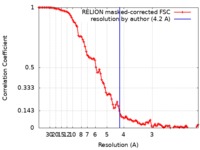
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Wang KT / Gourdon PE | |||||||||
| Funding support |  Denmark, 1 items Denmark, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2019 Journal: PLoS Biol / Year: 2019Title: Structure of the human ClC-1 chloride channel. Authors: Kaituo Wang / Sarah Spruce Preisler / Liying Zhang / Yanxiang Cui / Julie Winkel Missel / Christina Grønberg / Kamil Gotfryd / Erik Lindahl / Magnus Andersson / Kirstine Calloe / Pascal F ...Authors: Kaituo Wang / Sarah Spruce Preisler / Liying Zhang / Yanxiang Cui / Julie Winkel Missel / Christina Grønberg / Kamil Gotfryd / Erik Lindahl / Magnus Andersson / Kirstine Calloe / Pascal F Egea / Dan Arne Klaerke / Michael Pusch / Per Amstrup Pedersen / Z Hong Zhou / Pontus Gourdon /     Abstract: ClC-1 protein channels facilitate rapid passage of chloride ions across cellular membranes, thereby orchestrating skeletal muscle excitability. Malfunction of ClC-1 is associated with myotonia ...ClC-1 protein channels facilitate rapid passage of chloride ions across cellular membranes, thereby orchestrating skeletal muscle excitability. Malfunction of ClC-1 is associated with myotonia congenita, a disease impairing muscle relaxation. Here, we present the cryo-electron microscopy (cryo-EM) structure of human ClC-1, uncovering an architecture reminiscent of that of bovine ClC-K and CLC transporters. The chloride conducting pathway exhibits distinct features, including a central glutamate residue ("fast gate") known to confer voltage-dependence (a mechanistic feature not present in ClC-K), linked to a somewhat rearranged central tyrosine and a narrower aperture of the pore toward the extracellular vestibule. These characteristics agree with the lower chloride flux of ClC-1 compared with ClC-K and enable us to propose a model for chloride passage in voltage-dependent CLC channels. Comparison of structures derived from protein studied in different experimental conditions supports the notion that pH and adenine nucleotides regulate ClC-1 through interactions between the so-called cystathionine-β-synthase (CBS) domains and the intracellular vestibule ("slow gating"). The structure also provides a framework for analysis of mutations causing myotonia congenita and reveals a striking correlation between mutated residues and the phenotypic effect on voltage gating, opening avenues for rational design of therapies against ClC-1-related diseases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4657.map.gz emd_4657.map.gz | 84.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4657-v30.xml emd-4657-v30.xml emd-4657.xml emd-4657.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4657_fsc.xml emd_4657_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_4657.png emd_4657.png | 189.2 KB | ||
| Filedesc metadata |  emd-4657.cif.gz emd-4657.cif.gz | 6.7 KB | ||
| Others |  emd_4657_additional.map.gz emd_4657_additional.map.gz emd_4657_half_map_1.map.gz emd_4657_half_map_1.map.gz emd_4657_half_map_2.map.gz emd_4657_half_map_2.map.gz | 246.1 KB 70.2 MB 70.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4657 http://ftp.pdbj.org/pub/emdb/structures/EMD-4657 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4657 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4657 | HTTPS FTP |
-Related structure data
| Related structure data |  6qvuMC  4645C  4646C  4647C  4649C  6qv6C  6qvbC  6qvcC  6qvdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4657.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4657.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
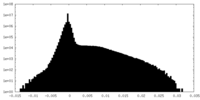
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_4657_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
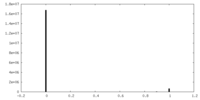
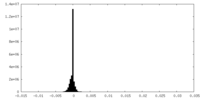
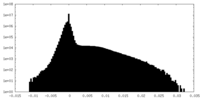
| Density Histograms |
-Half map: #2
| File | emd_4657_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
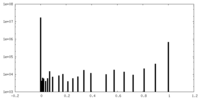
| Density Histograms |
-Half map: #1
| File | emd_4657_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
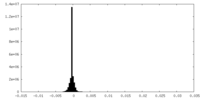
| Density Histograms |
- Sample components
Sample components
-Entire : human Chloride Channel protein 1
| Entire | Name: human Chloride Channel protein 1 |
|---|---|
| Components |
|
-Supramolecule #1: human Chloride Channel protein 1
| Supramolecule | Name: human Chloride Channel protein 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 109 KDa |
-Macromolecule #1: Chloride channel protein 1
| Macromolecule | Name: Chloride channel protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 108.733172 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEQSRSQQRG GEQSWWGSDP QYQYMPFEHC TSYGLPSENG GLQHRLRKDA GPRHNVHPTQ IYGHHKEQFS DREQDIGMPK KTGSSSTVD SKDEDHYSKC QDCIHRLGQV VRRKLGEDGI FLVLLGLLMA LVSWSMDYVS AKSLQAYKWS YAQMQPSLPL Q FLVWVTFP ...String: MEQSRSQQRG GEQSWWGSDP QYQYMPFEHC TSYGLPSENG GLQHRLRKDA GPRHNVHPTQ IYGHHKEQFS DREQDIGMPK KTGSSSTVD SKDEDHYSKC QDCIHRLGQV VRRKLGEDGI FLVLLGLLMA LVSWSMDYVS AKSLQAYKWS YAQMQPSLPL Q FLVWVTFP LVLILFSALF CHLISPQAVG SGIPEMKTIL RGVVLKEYLT MKAFVAKVVA LTAGLGSGIP VGKEGPFVHI AS ICAAVLS KFMSVFCGVY EQPYYYSDIL TVGCAVGVGC CFGTPLGGVL FSIEVTSTYF AVRNYWRGFF AATFSAFVFR VLA VWNKDA VTITALFRTN FRMDFPFDLK ELPAFAAIGI CCGLLGAVFV YLHRQVMLGV RKHKALSQFL AKHRLLYPGI VTFV IASFT FPPGMGQFMA GELMPREAIS TLFDNNTWVK HAGDPESLGQ SAVWIHPRVN VVIIIFLFFV MKFWMSIVAT TMPIP CGGF MPVFVLGAAF GRLVGEIMAM LFPDGILFDD IIYKILPGGY AVIGAAALTG AVSHTVSTAV ICFELTGQIA HILPMM VAV ILANMVAQSL QPSLYDSIIQ VKKLPYLPDL GWNQLSKYTI FVEDIMVRDV KFVSASYTYG ELRTLLQTTT VKTLPLV DS KDSMILLGSV ERSELQALLQ RHLCPERRLR AAQEMARKLS ELPYDGKARL AGEGLPGAPP GRPESFAFVD EDEDEDLS G KSELPPSLAL HPSTTAPLSP EEPNGPLPGH KQQPEAPEPA GQRPSIFQSL LHCLLGRARP TKKKTTQDST DLVDNMSPE EIEAWEQEQL SQPVCFDSCC IDQSPFQLVE QTTLHKTHTL FSLLGLHLAY VTSMGKLRGV LALEELQKAI EGHTKSGVQL RPPLASFRN TTSTRKSTGA PPSSAENWNL PEDRPGATGT GDVIAASPET PVPSPSPEPP LSLAPGKVEG ELEELELVES P GLEEELAD ILQGPSLRST DEEDEDELIL UniProtKB: Chloride channel protein 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Component - Concentration: 20.0 mM / Component - Name: BisTris / Details: 20mM BisTrisTris ph6.2 100mM NaCl 0.2mM TCEP |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 9.0 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)