[English] 日本語
 Yorodumi
Yorodumi- EMDB-43096: Structure of the E. coli clamp loader bound to the beta clamp in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the E. coli clamp loader bound to the beta clamp in an Initial-Binding conformation | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Bacterial Clamp Loader Complex / REPLICATION / TRANSFERASE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationDNA polymerase III, clamp loader complex / DNA clamp loader activity / DNA polymerase III complex / replisome / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / 3'-5' exonuclease activity / ribonucleoside triphosphate phosphatase activity / DNA-templated DNA replication / DNA replication ...DNA polymerase III, clamp loader complex / DNA clamp loader activity / DNA polymerase III complex / replisome / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / 3'-5' exonuclease activity / ribonucleoside triphosphate phosphatase activity / DNA-templated DNA replication / DNA replication / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / viral translational frameshifting / ATP hydrolysis activity / DNA binding / ATP binding / identical protein binding / metal ion binding / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
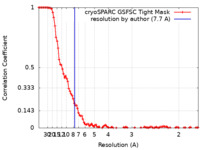
| Method | single particle reconstruction / cryo EM / Resolution: 7.7 Å | ||||||||||||
 Authors Authors | Landeck JT / Pajak J / Kelch BA | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2024 Journal: J Biol Chem / Year: 2024Title: Differences between bacteria and eukaryotes in clamp loader mechanism, a conserved process underlying DNA replication. Authors: Jacob T Landeck / Joshua Pajak / Emily K Norman / Emma L Sedivy / Brian A Kelch /  Abstract: Clamp loaders are pentameric ATPases that place circular sliding clamps onto DNA, where they function in DNA replication and genome integrity. The central activity of a clamp loader is the opening of ...Clamp loaders are pentameric ATPases that place circular sliding clamps onto DNA, where they function in DNA replication and genome integrity. The central activity of a clamp loader is the opening of the ring-shaped sliding clamp and the subsequent binding to primer-template (p/t)-junctions. The general architecture of clamp loaders is conserved across all life, suggesting that their mechanism is retained. Recent structural studies of the eukaryotic clamp loader replication factor C (RFC) revealed that it functions using a crab-claw mechanism, where clamp opening is coupled to a massive conformational change in the loader. Here we investigate the clamp loading mechanism of the Escherichia coli clamp loader at high resolution using cryo-electron microscopy. We find that the E. coli clamp loader opens the clamp using a crab-claw motion at a single pivot point, whereas the eukaryotic RFC loader uses motions distributed across the complex. Furthermore, we find clamp opening occurs in multiple steps, starting with a partly open state with a spiral conformation, and proceeding to a wide open clamp in a surprising planar geometry. Finally, our structures in the presence of p/t-junctions illustrate how the clamp closes around p/t-junctions and how the clamp loader initiates release from the loaded clamp. Our results reveal mechanistic distinctions in a macromolecular machine that is conserved across all domains of life. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43096.map.gz emd_43096.map.gz | 91.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43096-v30.xml emd-43096-v30.xml emd-43096.xml emd-43096.xml | 20.4 KB 20.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43096_fsc.xml emd_43096_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_43096.png emd_43096.png | 32 KB | ||
| Filedesc metadata |  emd-43096.cif.gz emd-43096.cif.gz | 6.9 KB | ||
| Others |  emd_43096_half_map_1.map.gz emd_43096_half_map_1.map.gz emd_43096_half_map_2.map.gz emd_43096_half_map_2.map.gz | 95.4 MB 95.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43096 http://ftp.pdbj.org/pub/emdb/structures/EMD-43096 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43096 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43096 | HTTPS FTP |
-Validation report
| Summary document |  emd_43096_validation.pdf.gz emd_43096_validation.pdf.gz | 948.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_43096_full_validation.pdf.gz emd_43096_full_validation.pdf.gz | 947.8 KB | Display | |
| Data in XML |  emd_43096_validation.xml.gz emd_43096_validation.xml.gz | 18.4 KB | Display | |
| Data in CIF |  emd_43096_validation.cif.gz emd_43096_validation.cif.gz | 23.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43096 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43096 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43096 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-43096 | HTTPS FTP |
-Related structure data
| Related structure data |  8vanMC  8valC  8vamC  8vapC  8vaqC  8varC  8vasC  8vatC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43096.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43096.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.87 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_43096_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_43096_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of the E. coli clamp loader bound to the beta clamp in ...
| Entire | Name: Structure of the E. coli clamp loader bound to the beta clamp in an Initial-Binding conformation |
|---|---|
| Components |
|
-Supramolecule #1: Structure of the E. coli clamp loader bound to the beta clamp in ...
| Supramolecule | Name: Structure of the E. coli clamp loader bound to the beta clamp in an Initial-Binding conformation type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: DNA polymerase III subunit delta
| Macromolecule | Name: DNA polymerase III subunit delta / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.745574 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIRLYPEQLR AQLNEGLRAA YLLLGNDPLL LQESQDAVRQ VAAAQGFEEH HTFSIDPNTD WNAIFSLCQA MSLFASRQTL LLLLPENGP NAAINEQLLT LTGLLHDDLL LIVRGNKLSK AQENAAWFTA LANRSVQVTC QTPEQAQLPR WVAARAKQLN L ELDDAANQ ...String: MIRLYPEQLR AQLNEGLRAA YLLLGNDPLL LQESQDAVRQ VAAAQGFEEH HTFSIDPNTD WNAIFSLCQA MSLFASRQTL LLLLPENGP NAAINEQLLT LTGLLHDDLL LIVRGNKLSK AQENAAWFTA LANRSVQVTC QTPEQAQLPR WVAARAKQLN L ELDDAANQ VLCYCYEGNL LALAQALERL SLLWPDGKLT LPRVEQAVND AAHFTPFHWV DALLMGKSKR ALHILQQLRL EG SEPVILL RTLQRELLLL VNLKRQSAHT PLRALFDKHR VWQNRRGMMG EALNRLSQTQ LRQAVQLLTR TELTLKQDYG QSV WAELEG LSLLLCHKPL ADVFIDG UniProtKB: DNA polymerase III subunit delta |
-Macromolecule #2: DNA polymerase III subunit tau
| Macromolecule | Name: DNA polymerase III subunit tau / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.803168 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPHMSYQVLA RKWRPQTFAD VVGQEHVLTA LANGLSLGRI HHAYLFSGTR GVGKTSIARL LAKGLNCETG ITATPCGVCD NCREIEQGR FVDLIEIDAA SRTKVEDTRD LLDNVQYAPA RGRFKVYLID EVHMLSRHSF NALLKTLEEP PEHVKFLLAT T DPQKLPVT ...String: GPHMSYQVLA RKWRPQTFAD VVGQEHVLTA LANGLSLGRI HHAYLFSGTR GVGKTSIARL LAKGLNCETG ITATPCGVCD NCREIEQGR FVDLIEIDAA SRTKVEDTRD LLDNVQYAPA RGRFKVYLID EVHMLSRHSF NALLKTLEEP PEHVKFLLAT T DPQKLPVT ILSRCLQFHL KALDVEQIRH QLEHILNEEH IAHEPRALQL LARAAEGSLR DALSLTDQAI ASGDGQVSTQ AV SAMLGTL DDDQALSLVE AMVEANGERV MALINEAAAR GIEWEALLVE MLGLLHRIAM VQLSPAALGN DMAAIELRMR ELA RTIPPT DIQLYYQTLL IGRKELPYAP DRRMGVEMTL LRALAFHPRM PLPEPEVPRQ UniProtKB: DNA polymerase III subunit tau |
-Macromolecule #3: DNA polymerase III subunit delta'
| Macromolecule | Name: DNA polymerase III subunit delta' / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 37.272801 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPHMRWYPWL RPDFEKLVAS YQAGRGHHAL LIQALPGMGD DALIYALSRY LLCQQPQGHK SCGHCRGCQL MQAGTHPDYY TLAPEKGKN TLGVDAVREV TEKLNEHARL GGAKVVWVTD AALLTDAAAN ALLKTLEEPP AETWFFLATR EPERLLATLR S RCRLHYLA ...String: GPHMRWYPWL RPDFEKLVAS YQAGRGHHAL LIQALPGMGD DALIYALSRY LLCQQPQGHK SCGHCRGCQL MQAGTHPDYY TLAPEKGKN TLGVDAVREV TEKLNEHARL GGAKVVWVTD AALLTDAAAN ALLKTLEEPP AETWFFLATR EPERLLATLR S RCRLHYLA PPPEQYAVTW LSREVTMSQD ALLAALRLSA GSPGAALALF QGDNWQARET LCQALAYSVP SGDWYSLLAA LN HEQAPAR LHWLATLLMD ALKRHHGAAQ VTNVDVPGLV AELANHLSPS RLQAILGDVC HIREQLMSVT GINRELLITD LLL RIEHYL QPGVVLPVPH L UniProtKB: DNA polymerase III subunit delta' |
-Macromolecule #4: Beta sliding clamp
| Macromolecule | Name: Beta sliding clamp / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.922816 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPHMKFTVER EHLLKPLQQV SGPLGGRPTL PILGNLLLQV ADGTLSLTGT DLEMEMVARV ALVQPHEPGA TTVPARKFFD ICRGLPEGA EIAVQLEGER MLVRSGRSRF SLSTLPAADF PNLDDWQSEV EFTLPQATMK RLIEATQFSM AHQDVRYYLN G MLFETEGE ...String: GPHMKFTVER EHLLKPLQQV SGPLGGRPTL PILGNLLLQV ADGTLSLTGT DLEMEMVARV ALVQPHEPGA TTVPARKFFD ICRGLPEGA EIAVQLEGER MLVRSGRSRF SLSTLPAADF PNLDDWQSEV EFTLPQATMK RLIEATQFSM AHQDVRYYLN G MLFETEGE ELRTVATDGH RLAVCSMPIG QSLPSHSVIV PRKGVIELMR MLDGGDNPLR VQIGSNNIRA HVGDFIFTSK LV DGRFPDY RRVLPKNPDK HLEAGCDLLK QAFARAAILS NEKFRGVRLY VSENQLKITA NNPEQEEAEE ILDVTYSGAE MEI GFNVSY VLDVLNALKC ENVRMMLTDS VSSVQIEDAA SQSAAYVVMP MRL UniProtKB: Beta sliding clamp |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 3664 / Average electron dose: 42.8 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.3000000000000003 µm / Nominal defocus min: 1.1 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)