+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of PI3Kalpha | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PI3Kalapha / isoform selective / structure-based drug design. / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationperinuclear endoplasmic reticulum membrane / response to muscle inactivity / regulation of toll-like receptor 4 signaling pathway / negative regulation of actin filament depolymerization / phosphatidylinositol kinase activity / response to L-leucine / regulation of actin filament organization / response to butyrate / phosphatidylinositol 3-kinase regulator activity / IRS-mediated signalling ...perinuclear endoplasmic reticulum membrane / response to muscle inactivity / regulation of toll-like receptor 4 signaling pathway / negative regulation of actin filament depolymerization / phosphatidylinositol kinase activity / response to L-leucine / regulation of actin filament organization / response to butyrate / phosphatidylinositol 3-kinase regulator activity / IRS-mediated signalling / positive regulation of focal adhesion disassembly / cellular response to hydrostatic pressure / phosphatidylinositol 3-kinase activator activity / autosome genomic imprinting / interleukin-18-mediated signaling pathway / PI3K events in ERBB4 signaling / myeloid leukocyte migration / 1-phosphatidylinositol-3-kinase regulator activity / phosphatidylinositol 3-kinase regulatory subunit binding / neurotrophin TRKA receptor binding / Activated NTRK2 signals through PI3K / positive regulation of endoplasmic reticulum unfolded protein response / positive regulation of protein localization to membrane / Activated NTRK3 signals through PI3K / negative regulation of fibroblast apoptotic process / cis-Golgi network / ErbB-3 class receptor binding / kinase activator activity / phosphatidylinositol 3-kinase complex, class IB / vasculature development / transmembrane receptor protein tyrosine kinase adaptor activity / regulation of cellular respiration / RHOF GTPase cycle / Signaling by cytosolic FGFR1 fusion mutants / RHOD GTPase cycle / cardiac muscle cell contraction / phosphatidylinositol 3-kinase complex, class IA / phosphatidylinositol 3-kinase complex / enzyme-substrate adaptor activity / Nephrin family interactions / anoikis / Signaling by LTK in cancer / Costimulation by the CD28 family / RND1 GTPase cycle / Signaling by LTK / 1-phosphatidylinositol-4-phosphate 3-kinase activity / 1-phosphatidylinositol-4,5-bisphosphate 3-kinase activity / relaxation of cardiac muscle / MET activates PI3K/AKT signaling / positive regulation of leukocyte migration / PI3K/AKT activation / RND2 GTPase cycle / phosphatidylinositol-4,5-bisphosphate 3-kinase / positive regulation of filopodium assembly / RND3 GTPase cycle / growth hormone receptor signaling pathway / vascular endothelial growth factor signaling pathway / negative regulation of stress fiber assembly / phosphatidylinositol 3-kinase / insulin binding / phosphatidylinositol-3-phosphate biosynthetic process / natural killer cell mediated cytotoxicity / 1-phosphatidylinositol-3-kinase activity / RHOV GTPase cycle / negative regulation of cell-matrix adhesion / Signaling by ALK / negative regulation of macroautophagy / RHOB GTPase cycle / GP1b-IX-V activation signalling / PI-3K cascade:FGFR2 / PI-3K cascade:FGFR3 / Erythropoietin activates Phosphoinositide-3-kinase (PI3K) / protein kinase activator activity / PI-3K cascade:FGFR4 / response to dexamethasone / PI-3K cascade:FGFR1 / RHOC GTPase cycle / RHOJ GTPase cycle / negative regulation of osteoclast differentiation / intracellular glucose homeostasis / Synthesis of PIPs at the plasma membrane / phosphatidylinositol-mediated signaling / phosphatidylinositol phosphate biosynthetic process / CD28 dependent PI3K/Akt signaling / RHOU GTPase cycle / CDC42 GTPase cycle / PI3K events in ERBB2 signaling / negative regulation of anoikis / PI3K Cascade / RET signaling / intercalated disc / insulin receptor substrate binding / Interleukin-3, Interleukin-5 and GM-CSF signaling / T cell differentiation / RHOG GTPase cycle / regulation of multicellular organism growth / extrinsic apoptotic signaling pathway via death domain receptors / endothelial cell migration / positive regulation of TOR signaling / RHOA GTPase cycle Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
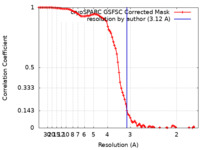
| Method | single particle reconstruction / cryo EM / Resolution: 3.12 Å | |||||||||
 Authors Authors | Valverde R / Shi H / Holliday M / Sun M | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Cancer Discov / Year: 2024 Journal: Cancer Discov / Year: 2024Title: Discovery and Clinical Proof-of-Concept of RLY-2608, a First-in-Class Mutant-Selective Allosteric PI3Kα Inhibitor That Decouples Antitumor Activity from Hyperinsulinemia. Authors: Andreas Varkaris / Ermira Pazolli / Hakan Gunaydin / Qi Wang / Levi Pierce / Alessandro A Boezio / Artemisa Bulku / Lucian DiPietro / Cary Fridrich / Adam Frost / Fabrizio Giordanetto / ...Authors: Andreas Varkaris / Ermira Pazolli / Hakan Gunaydin / Qi Wang / Levi Pierce / Alessandro A Boezio / Artemisa Bulku / Lucian DiPietro / Cary Fridrich / Adam Frost / Fabrizio Giordanetto / Erika P Hamilton / Katherine Harris / Michael Holliday / Tamieka L Hunter / Amanda Iskandar / Yongli Ji / Alexandre Larivée / Jonathan R LaRochelle / André Lescarbeau / Fabien Llambi / Brenda Lormil / Mary M Mader / Brenton G Mar / Iain Martin / Thomas H McLean / Klaus Michelsen / Yakov Pechersky / Erika Puente-Poushnejad / Kevin Raynor / Dipali Rogala / Ramin Samadani / Alison M Schram / Kelley Shortsleeves / Sweta Swaminathan / Shahein Tajmir / Gege Tan / Yong Tang / Roberto Valverde / Bryan Wehrenberg / Jeremy Wilbur / Bret R Williams / Hongtao Zeng / Hanmo Zhang / W Patrick Walters / Beni B Wolf / David E Shaw / Donald A Bergstrom / James Watters / James S Fraser / Pascal D Fortin / D Randal Kipp /   Abstract: PIK3CA (PI3Kα) is a lipid kinase commonly mutated in cancer, including ∼40% of hormone receptor-positive breast cancer. The most frequently observed mutants occur in the kinase and helical domains. ...PIK3CA (PI3Kα) is a lipid kinase commonly mutated in cancer, including ∼40% of hormone receptor-positive breast cancer. The most frequently observed mutants occur in the kinase and helical domains. Orthosteric PI3Kα inhibitors suffer from poor selectivity leading to undesirable side effects, most prominently hyperglycemia due to inhibition of wild-type (WT) PI3Kα. Here, we used molecular dynamics simulations and cryo-electron microscopy to identify an allosteric network that provides an explanation for how mutations favor PI3Kα activation. A DNA-encoded library screen leveraging electron microscopy-optimized constructs, differential enrichment, and an orthosteric-blocking compound led to the identification of RLY-2608, a first-in-class allosteric mutant-selective inhibitor of PI3Kα. RLY-2608 inhibited tumor growth in PIK3CA-mutant xenograft models with minimal impact on insulin, a marker of dysregulated glucose homeostasis. RLY-2608 elicited objective tumor responses in two patients diagnosed with advanced hormone receptor-positive breast cancer with kinase or helical domain PIK3CA mutations, with no observed WT PI3Kα-related toxicities. SIGNIFICANCE: Treatments for PIK3CA-mutant cancers are limited by toxicities associated with the inhibition of WT PI3Kα. Molecular dynamics, cryo-electron microscopy, and DNA-encoded libraries were ...SIGNIFICANCE: Treatments for PIK3CA-mutant cancers are limited by toxicities associated with the inhibition of WT PI3Kα. Molecular dynamics, cryo-electron microscopy, and DNA-encoded libraries were used to develop RLY-2608, a first-in-class inhibitor that demonstrates mutant selectivity in patients. This marks the advance of clinical mutant-selective inhibition that overcomes limitations of orthosteric PI3Kα inhibitors. See related commentary by Gong and Vanhaesebroeck, p. 204 . See related article by Varkaris et al., p. 227 . This article is featured in Selected Articles from This Issue, p. 201. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41617.map.gz emd_41617.map.gz | 3.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41617-v30.xml emd-41617-v30.xml emd-41617.xml emd-41617.xml | 25.6 KB 25.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41617_fsc.xml emd_41617_fsc.xml | 9.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_41617.png emd_41617.png | 69.8 KB | ||
| Filedesc metadata |  emd-41617.cif.gz emd-41617.cif.gz | 7.7 KB | ||
| Others |  emd_41617_additional_1.map.gz emd_41617_additional_1.map.gz emd_41617_half_map_1.map.gz emd_41617_half_map_1.map.gz emd_41617_half_map_2.map.gz emd_41617_half_map_2.map.gz | 81.7 MB 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41617 http://ftp.pdbj.org/pub/emdb/structures/EMD-41617 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41617 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41617 | HTTPS FTP |
-Validation report
| Summary document |  emd_41617_validation.pdf.gz emd_41617_validation.pdf.gz | 733.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41617_full_validation.pdf.gz emd_41617_full_validation.pdf.gz | 733.3 KB | Display | |
| Data in XML |  emd_41617_validation.xml.gz emd_41617_validation.xml.gz | 17.8 KB | Display | |
| Data in CIF |  emd_41617_validation.cif.gz emd_41617_validation.cif.gz | 23.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41617 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41617 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41617 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41617 | HTTPS FTP |
-Related structure data
| Related structure data |  8tu6MC  8ts7C  8ts8C  8ts9C  8tsaC  8tsbC  8tscC  8tsdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41617.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41617.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.819 Å | ||||||||||||||||||||||||||||||||||||
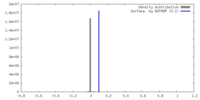
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_41617_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
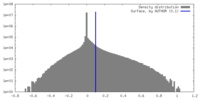
| Density Histograms |
-Half map: #2
| File | emd_41617_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
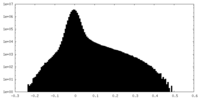
| Density Histograms |
-Half map: #1
| File | emd_41617_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PI3Kalpha
| Entire | Name: PI3Kalpha |
|---|---|
| Components |
|
-Supramolecule #1: PI3Kalpha
| Supramolecule | Name: PI3Kalpha / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 850 KDa |
-Supramolecule #2: p110
| Supramolecule | Name: p110 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: p85
| Supramolecule | Name: p85 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit ...
| Macromolecule | Name: Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 126.301992 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MHHHHHHGSL EVLFQGPPPR PSSGELWGIH LMPPRILVEC LLPNGMIVTL ECLREATLIT IKHELFKEAR KYPLHQLLQD ESSYIFVSV TQEAEREEFF DETRRLCDLR LFQPFLKVIE PVGNREEKIL NREIGFAIGM PVCEFDMVKD PEVQDFRRNI L NVCKEAVD ...String: MHHHHHHGSL EVLFQGPPPR PSSGELWGIH LMPPRILVEC LLPNGMIVTL ECLREATLIT IKHELFKEAR KYPLHQLLQD ESSYIFVSV TQEAEREEFF DETRRLCDLR LFQPFLKVIE PVGNREEKIL NREIGFAIGM PVCEFDMVKD PEVQDFRRNI L NVCKEAVD LRDLNSPHSR AMYVYPPNVE SSPELPKHIY NKLDKGQIIV VIWVIVSPNN DKQKYTLKIN HDCVPEQVIA EA IRKKTRS MLLSSEQLKL CVLEYQGKYI LKVCGCDEYF LEKYPLSQYK YIRSCIMLGR MPNLMLMAKE SLYSQLPMDC FTM PSYSRR ISTATPYMNG ETSTKSLWVI NSALRIKILC ATYVNVNIRD IDKIYVRTGI YHGGEPLCDN VNTQRVPCSN PRWN EWLNY DIYIPDLPRA ARLCLSICSV KGRKGAKEEH CPLAWGNINL FDYTDTLVSG KMALNLWPVP HGLEDLLNPI GVTGS NPNK ETPCLELEFD WFSSVVKFPD MSVIEEHANW SVSREAGFSY SHAGLSNRLA RDNELRENDK EQLKAISTRD PLSEIT EQE KDFLWSHRHY CVTIPEILPK LLLSVKWNSR DEVAQMYCLV KDWPPIKPEQ AMELLDCNYP DPMVRGFAVR CLEKYLT DD KLSQYLIQLV QVLKYEQYLD NLLVRFLLKK ALTNQRIGHF FFWHLKSEMH NKTVSQRFGL LLESYCRACG MYLKHLNR Q VEAMEKLINL TDILKQEKKD ETQKVQMKFL VEQMRRPDFM DALQGFLSPL NPAHQLGNLR LEECRIMSSA KRPLWLNWE NPDIMSELLF QNNEIIFKNG DDLRQDMLTL QIIRIMENIW QNQGLDLRML PYGCLSIGDC VGLIEVVRNS HTIMQIQCKG GLKGALQFN SHTLHQWLKD KNKGEIYDAA IDLFTRSCAG YCVATFILGI GDRHNSNIMV KDDGQLFHID FGHFLDHKKK K FGYKRERV PFVLTQDFLI VISKGAQECT KTREFERFQE MCYKAYLAIR QHANLFINLF SMMLGSGMPE LQSFDDIAYI RK TLALDKT EQEALEYFMK QMNDAHHGGW TTKMDWIFHT IKQHALN UniProtKB: Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform |
-Macromolecule #2: Phosphatidylinositol 3-kinase regulatory subunit alpha
| Macromolecule | Name: Phosphatidylinositol 3-kinase regulatory subunit alpha type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 83.710281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSAEGYQYRA LYDYKKEREE DIDLHLGDIL TVNKGSLVAL GFSDGQEARP EEIGWLNGYN ETTGERGDFP GTYVEYIGRK KISPPTPKP RPPRPLPVAP GSSKTEADVE QQALTLPDLA EQFAPPDIAP PLLIKLVEAI EKKGLECSTL YRTQSSSNLA E LRQLLDCD ...String: MSAEGYQYRA LYDYKKEREE DIDLHLGDIL TVNKGSLVAL GFSDGQEARP EEIGWLNGYN ETTGERGDFP GTYVEYIGRK KISPPTPKP RPPRPLPVAP GSSKTEADVE QQALTLPDLA EQFAPPDIAP PLLIKLVEAI EKKGLECSTL YRTQSSSNLA E LRQLLDCD TPSVDLEMID VHVLADAFKR YLLDLPNPVI PAAVYSEMIS LAPEVQSSEE YIQLLKKLIR SPSIPHQYWL TL QYLLKHF FKLSQTSSKN LLNARVLSEI FSPMLFRFSA ASSDNTENLI KVIEILISTE WNERQPAPAL PPKPPKPTTV ANN GMNNNM SLQDAEWYWG DISREEVNEK LRDTADGTFL VRDASTKMHG DYTLTLRKGG NNKLIKIFHR DGKYGFSDPL TFSS VVELI NHYRNESLAQ YNPKLDVKLL YPVSKYQQDQ VVKEDNIEAV GKKLHEYNTQ FQEKSREYDR LYEEYTRTSQ EIQMK RTAI EAFNETIKIF EEQCQTQERY SKEYIEKFKR EGNEKEIQRI MHNYDKLKSR ISEIIDSRRR LEEDLKKQAA EYREID KRM NSIKPDLIQL RKTRDQYLMW LTQKGVRQKK LNEWLGNENT EDQYSLVEDD EDLPHHDEKT WNVGSSNRNK AENLLRG KR DGTFLVRESS KQGCYACSVV VDGEVKHCVI NKTATGYGFA EPYNLYSSLK ELVLHYQHTS LVQHNDSLNV TLAYPVYA Q QRR UniProtKB: Phosphatidylinositol 3-kinase regulatory subunit alpha |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.35 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Support film - Film thickness: 50 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. |
| Vitrification | Cryogen name: ETHANE |
| Details | The sample was purified as a heterodimer and was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K3 (6k x 4k) / #0 - Detector mode: COUNTING / #0 - Number grids imaged: 3 / #0 - Number real images: 6866 / #0 - Average exposure time: 2.68 sec. / #0 - Average electron dose: 60.0 e/Å2 #0 - Details: Grids were prepared with 0.02% CTAB. Images were collected in movie mode at 23 frames per second. #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K3 (6k x 4k) / #1 - Number grids imaged: 1 / #1 - Number real images: 2058 / #1 - Average exposure time: 2.68 sec. / #1 - Average electron dose: 60.0 e/Å2 #1 - Details: Images were collected in movie mode at 23 frames per second. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)