[English] 日本語
 Yorodumi
Yorodumi- EMDB-40959: Cryo-EM structure of human TRPV4 ankyrin repeat domain in complex... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
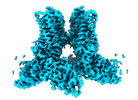
| Title | Cryo-EM structure of human TRPV4 ankyrin repeat domain in complex with GTPase RhoA | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | transient receptor potential V family member 4 / TRP / channel / TRPV4 / TRP channels / membrane protein / GTPase / transforming protein RhoA / RhoA / ankyrin repeat domain / GDP / guanosine diphosphate | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationstretch-activated, monoatomic cation-selective, calcium channel activity / blood vessel endothelial cell delamination / osmosensor activity / vasopressin secretion / positive regulation of striated muscle contraction / calcium ion import into cytosol / positive regulation of macrophage inflammatory protein 1 alpha production / negative regulation of brown fat cell differentiation / positive regulation of microtubule depolymerization / hyperosmotic salinity response ...stretch-activated, monoatomic cation-selective, calcium channel activity / blood vessel endothelial cell delamination / osmosensor activity / vasopressin secretion / positive regulation of striated muscle contraction / calcium ion import into cytosol / positive regulation of macrophage inflammatory protein 1 alpha production / negative regulation of brown fat cell differentiation / positive regulation of microtubule depolymerization / hyperosmotic salinity response / cortical microtubule organization / regulation of response to osmotic stress / positive regulation of chemokine (C-X-C motif) ligand 1 production / positive regulation of chemokine (C-C motif) ligand 5 production / cartilage development involved in endochondral bone morphogenesis / aortic valve formation / alpha-beta T cell lineage commitment / mitotic cleavage furrow formation / bone trabecula morphogenesis / positive regulation of lipase activity / endothelial tube lumen extension / skeletal muscle satellite cell migration / positive regulation of vascular associated smooth muscle contraction / angiotensin-mediated vasoconstriction involved in regulation of systemic arterial blood pressure / SLIT2:ROBO1 increases RHOA activity / RHO GTPases Activate Rhotekin and Rhophilins / Roundabout signaling pathway / cellular hypotonic response / negative regulation of intracellular steroid hormone receptor signaling pathway / Axonal growth inhibition (RHOA activation) / Axonal growth stimulation / cellular hypotonic salinity response / regulation of neural precursor cell proliferation / cleavage furrow formation / regulation of modification of postsynaptic actin cytoskeleton / regulation of osteoblast proliferation / multicellular organismal-level water homeostasis / osmosensory signaling pathway / forebrain radial glial cell differentiation / cell junction assembly / apical junction assembly / regulation of systemic arterial blood pressure by endothelin / cerebral cortex cell migration / positive regulation of vascular permeability / negative regulation of cell migration involved in sprouting angiogenesis / cellular response to chemokine / beta selection / establishment of epithelial cell apical/basal polarity / negative regulation of cell size / regulation of modification of postsynaptic structure / RHO GTPases Activate ROCKs / negative regulation of oxidative phosphorylation / negative regulation of motor neuron apoptotic process / ERBB2 Regulates Cell Motility / cellular response to osmotic stress / RHO GTPases activate CIT / cell volume homeostasis / positive regulation of monocyte chemotactic protein-1 production / Sema4D induced cell migration and growth-cone collapse / calcium ion import / PCP/CE pathway / RHO GTPases activate KTN1 / apolipoprotein A-I-mediated signaling pathway / positive regulation of podosome assembly / cell-cell junction assembly / negative regulation of cell-substrate adhesion / positive regulation of alpha-beta T cell differentiation / TRP channels / ossification involved in bone maturation / odontogenesis / Wnt signaling pathway, planar cell polarity pathway / Sema4D mediated inhibition of cell attachment and migration / motor neuron apoptotic process / positive regulation of leukocyte adhesion to vascular endothelial cell / PI3K/AKT activation / wound healing, spreading of cells / apical junction complex / regulation of focal adhesion assembly / negative chemotaxis / regulation of aerobic respiration / cortical actin cytoskeleton / myosin binding / EPHA-mediated growth cone collapse / positive regulation of macrophage chemotaxis / stress fiber assembly / regulation of neuron projection development / RHOC GTPase cycle / beta-tubulin binding / cytoplasmic microtubule / diet induced thermogenesis / positive regulation of cytokinesis / androgen receptor signaling pathway / cellular response to cytokine stimulus / microtubule polymerization / cleavage furrow / bioluminescence / semaphorin-plexin signaling pathway / Rho protein signal transduction / ficolin-1-rich granule membrane / mitotic spindle assembly Similarity search - Function | |||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human cytomegalovirus Human cytomegalovirus | |||||||||||||||||||||
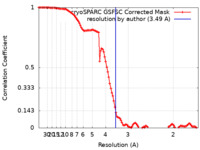
| Method | single particle reconstruction / cryo EM / Resolution: 3.49 Å | |||||||||||||||||||||
 Authors Authors | Nadezhdin KD / Talyzina IA / Neuberger A / Sobolevsky AI | |||||||||||||||||||||
| Funding support |  United States, United States,  Germany, 6 items Germany, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structure of human TRPV4 in complex with GTPase RhoA. Authors: Kirill D Nadezhdin / Irina A Talyzina / Aravind Parthasarathy / Arthur Neuberger / David X Zhang / Alexander I Sobolevsky /  Abstract: Transient receptor potential (TRP) channel TRPV4 is a polymodal cellular sensor that responds to moderate heat, cell swelling, shear stress, and small-molecule ligands. It is involved in ...Transient receptor potential (TRP) channel TRPV4 is a polymodal cellular sensor that responds to moderate heat, cell swelling, shear stress, and small-molecule ligands. It is involved in thermogenesis, regulation of vascular tone, bone homeostasis, renal and pulmonary functions. TRPV4 is implicated in neuromuscular and skeletal disorders, pulmonary edema, and cancers, and represents an important drug target. The cytoskeletal remodeling GTPase RhoA has been shown to suppress TRPV4 activity. Here, we present a structure of the human TRPV4-RhoA complex that shows RhoA interaction with the membrane-facing surface of the TRPV4 ankyrin repeat domains. The contact interface reveals residues that are mutated in neuropathies, providing an insight into the disease pathogenesis. We also identify the binding sites of the TRPV4 agonist 4α-PDD and the inhibitor HC-067047 at the base of the S1-S4 bundle, and show that agonist binding leads to pore opening, while channel inhibition involves a π-to-α transition in the pore-forming helix S6. Our structures elucidate the interaction interface between hTRPV4 and RhoA, as well as residues at this interface that are involved in TRPV4 disease-causing mutations. They shed light on TRPV4 activation and inhibition and provide a template for the design of future therapeutics for treatment of TRPV4-related diseases. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40959.map.gz emd_40959.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40959-v30.xml emd-40959-v30.xml emd-40959.xml emd-40959.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40959_fsc.xml emd_40959_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_40959.png emd_40959.png | 89.7 KB | ||
| Others |  emd_40959_half_map_1.map.gz emd_40959_half_map_1.map.gz emd_40959_half_map_2.map.gz emd_40959_half_map_2.map.gz | 115.8 MB 115.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40959 http://ftp.pdbj.org/pub/emdb/structures/EMD-40959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40959 | HTTPS FTP |
-Related structure data
| Related structure data |  8t1cMC  8t1bC  8t1dC  8t1eC  8t1fC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40959.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40959.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_40959_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_40959_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human TRPV4 ankyrin repeat domain in complex with GTPase RhoA
| Entire | Name: human TRPV4 ankyrin repeat domain in complex with GTPase RhoA |
|---|---|
| Components |
|
-Supramolecule #1: human TRPV4 ankyrin repeat domain in complex with GTPase RhoA
| Supramolecule | Name: human TRPV4 ankyrin repeat domain in complex with GTPase RhoA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 510 KDa |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 4-...
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 4-Enhanced green fluorescent protein chimera type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human cytomegalovirus Human cytomegalovirus |
| Molecular weight | Theoretical: 127.717938 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADSSEGPRA GPGEVAELPG DESGTPGGEA FPLSSLANLF EGEDGSLSPS PADASRPAGP GDGRPNLRMK FQGAFRKGVP NPIDLLEST LYESSVVPGP KKAPMDSLFD YGTYRHHSSD NKRWRKKIIE KQPQSPKAPA PQPPPILKVF NRPILFDIVS R GSTADLDG ...String: MADSSEGPRA GPGEVAELPG DESGTPGGEA FPLSSLANLF EGEDGSLSPS PADASRPAGP GDGRPNLRMK FQGAFRKGVP NPIDLLEST LYESSVVPGP KKAPMDSLFD YGTYRHHSSD NKRWRKKIIE KQPQSPKAPA PQPPPILKVF NRPILFDIVS R GSTADLDG LLPFLLTHKK RLTDEEFREP STGKTCLPKA LLNLSNGRND TIPVLLDIAE RTGNMREFIN SPFRDIYYRG QT ALHIAIE RRCKHYVELL VAQGADVHAQ ARGRFFQPKD EGGYFYFGEL PLSLAACTNQ PHIVNYLTEN PHKKADMRRQ DSR GNTVLH ALVAIADNTR ENTKFVTKMY DLLLLKCARL FPDSNLEAVL NNDGLSPLMM AAKTGKIGIF QHIIRREVTD EDTR HLSRK FKDWAYGPVY SSLYDLSSLD TCGEEASVLE ILVYNSKIEN RHEMLAVEPI NELLRDKWRK FGAVSFYINV VSYLC AMVI FTLTAYYQPL EGTPPYPYRT TVDYLRLAGE VITLFTGVLF FFTNIKDLFM KKCPGVNSLF IDGSFQLLYF IYSVLV IVS AALYLAGIEA YLAVMVFALV LGWMNALYFT RGLKLTGTYS IMIQKILFKD LFRFLLVYLL FMIGYASALV SLLNPCA NM KVCNEDQTNC TVPTYPSCRD SETFSTFLLD LFKLTIGMGD LEMLSSTKYP VVFIILLVTY IILTFVLLLN MLIALMGE T VGQVSKESKH IWKLQWATTI LDIERSFPVF LRKAFRSGEM VTVGKSSDGT PDRRWCFRVD EVNWSHWNQN LGIINEDPG KNETYQYYGF SHTVGRLRRD RWSSVVPRVV ELNKNSNPDE VVVPLDSMGN PRCDGHQQGY PRKWRTDDAP LLVPRGSAAA AVSKGEELF TGVVPILVEL DGDVNGHKFS VSGEGEGDAT YGKLTLKFIC TTGKLPVPWP TLVTTLTYGV QCFSRYPDHM K QHDFFKSA MPEGYVQERT IFFKDDGNYK TRAEVKFEGD TLVNRIELKG IDFKEDGNIL GHKLEYNYNS HNVYIMADKQ KN GIKVNFK IRHNIEDGSV QLADHYQQNT PIGDGPVLLP DNHYLSTQSK LSKDPNEKRD HMVLLEFVTA AGITLGMDEL YKS GLRSWS HPQFEK |
-Macromolecule #2: Transforming protein RhoA
| Macromolecule | Name: Transforming protein RhoA / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: small monomeric GTPase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.799158 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAAIRKKLVI VGDGACGKTC LLIVFSKDQF PEVYVPTVFE NYVADIEVDG KQVELALWDT AGQEDYDRLR PLSYPDTDVI LMCFSIDSP DSLENIPEKW TPEVKHFCPN VPIILVGNKK DLRNDEHTRR ELAKMKQEPV KPEEGRDMAN RIGAFGYMEC S AKTKDGVR ...String: MAAIRKKLVI VGDGACGKTC LLIVFSKDQF PEVYVPTVFE NYVADIEVDG KQVELALWDT AGQEDYDRLR PLSYPDTDVI LMCFSIDSP DSLENIPEKW TPEVKHFCPN VPIILVGNKK DLRNDEHTRR ELAKMKQEPV KPEEGRDMAN RIGAFGYMEC S AKTKDGVR EVFEMATRAA LQARRGKKKS GCLVL |
-Macromolecule #3: GUANOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 1 / Formula: GDP |
|---|---|
| Molecular weight | Theoretical: 443.201 Da |
| Chemical component information |  ChemComp-GDP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | human TRPV4 |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 5456 / Average exposure time: 2.5 sec. / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8t1c: |
 Movie
Movie Controller
Controller




















 Z
Z Y
Y X
X