+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3nx5 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
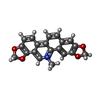
| Title | The crystal structure of Sanguinarine bound to DNA d(CGTACG) | ||||||||||||||||||
 Components Components | 5'-D(* Keywords KeywordsDNA / DRUG-DNA COMPLEX / DOUBLE HELIX | Function / homology | Chem-SAU / DNA |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 2.31 Å molecular replacement / Resolution: 2.31 Å  Authors AuthorsFerraroni, M. / Bazzicalupi, C. / Gratteri, P. / Bilia, A.R. |  Citation Citation Journal: Chem.Commun.(Camb.) / Year: 2011 Journal: Chem.Commun.(Camb.) / Year: 2011Title: X-Ray diffraction analyses of the natural isoquinoline alkaloids Berberine and Sanguinarine complexed with double helix DNA d(CGTACG) Authors: Ferraroni, M. / Bazzicalupi, C. / Bilia, A.R. / Gratteri, P. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3nx5.cif.gz 3nx5.cif.gz | 23.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3nx5.ent.gz pdb3nx5.ent.gz | 14.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3nx5.json.gz 3nx5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/nx/3nx5 https://data.pdbj.org/pub/pdb/validation_reports/nx/3nx5 ftp://data.pdbj.org/pub/pdb/validation_reports/nx/3nx5 ftp://data.pdbj.org/pub/pdb/validation_reports/nx/3nx5 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3np6SC S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 1809.217 Da / Num. of mol.: 4 / Source method: obtained synthetically / Details: Synthetic DNA #2: Chemical | ChemComp-SAU / | #3: Chemical | ChemComp-CA / | #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.14 Å3/Da / Density % sol: 42.42 % |
|---|---|
| Crystal grow | Temperature: 296 K / Method: vapor diffusion / pH: 6.5 Details: MPD, MgCl2, NaCl, pH 6.5, vapor diffusion, temperature 296K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.9763 Å / Beamline: ID29 / Wavelength: 0.9763 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Feb 8, 2010 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: Si(111) monochromator / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9763 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection twin |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.29→39.69 Å / Num. obs: 3006 / % possible obs: 96.3 % / Observed criterion σ(I): -3 / Redundancy: 4.9 % / Biso Wilson estimate: 47.698 Å2 / Rmerge(I) obs: 0.051 / Rsym value: 0.051 / Net I/σ(I): 20.99 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR |
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 3NP6 Resolution: 2.31→39.69 Å / Cor.coef. Fo:Fc: 0.921 / Cor.coef. Fo:Fc free: 0.892 / WRfactor Rfree: 0.3248 / WRfactor Rwork: 0.2852 / Occupancy max: 1 / Occupancy min: 1 / FOM work R set: 0.5756 / SU B: 14.156 / SU ML: 0.294 / SU R Cruickshank DPI: 0.0981 / SU Rfree: 0.0596 / Cross valid method: THROUGHOUT / ESU R Free: 0.06 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: U VALUES: REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 60.94 Å2 / Biso mean: 33.9258 Å2 / Biso min: 13.33 Å2
| |||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.31→39.69 Å
| |||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.305→2.365 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller









 PDBj
PDBj