[English] 日本語
 Yorodumi
Yorodumi- EMDB-3805: Cryo-EM structure of jasplakinolide-stabilized malaria parasite F... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3805 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of jasplakinolide-stabilized malaria parasite F-actin at near-atomic resolution | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | F-actin / Plasmodium / malaria parasite / cytoskeleton / cryo-EM / JAS / Jasplakinolide / filament / glideosome / gliding motility / thin filament / structural protein | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated actin polymerization-dependent cell-to-cell migration in host / actin filament / structural constituent of cytoskeleton / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / hydrolase activity / ATP binding / nucleus / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||||||||
 Authors Authors | Pospich S / Kumpula E-P | |||||||||||||||
| Funding support |  Germany, Germany,  Finland, 4 items Finland, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2017 Journal: Proc Natl Acad Sci U S A / Year: 2017Title: Near-atomic structure of jasplakinolide-stabilized malaria parasite F-actin reveals the structural basis of filament instability. Authors: Sabrina Pospich / Esa-Pekka Kumpula / Julian von der Ecken / Juha Vahokoski / Inari Kursula / Stefan Raunser /    Abstract: During their life cycle, apicomplexan parasites, such as the malaria parasite , use actomyosin-driven gliding motility to move and invade host cells. For this process, actin filament length and ...During their life cycle, apicomplexan parasites, such as the malaria parasite , use actomyosin-driven gliding motility to move and invade host cells. For this process, actin filament length and stability are temporally and spatially controlled. In contrast to canonical actin, actin 1 (Act1) does not readily polymerize into long, stable filaments. The structural basis of filament instability, which plays a pivotal role in host cell invasion, and thus infectivity, is poorly understood, largely because high-resolution structures of Act1 filaments were missing. Here, we report the near-atomic structure of jasplakinolide (JAS)-stabilized Act1 filaments determined by electron cryomicroscopy. The general filament architecture is similar to that of mammalian F-actin. The high resolution of the structure allowed us to identify small but important differences at inter- and intrastrand contact sites, explaining the inherent instability of apicomplexan actin filaments. JAS binds at regular intervals inside the filament to three adjacent actin subunits, reinforcing filament stability by hydrophobic interactions. Our study reveals the high-resolution structure of a small molecule bound to F-actin, highlighting the potential of electron cryomicroscopy for structure-based drug design. Furthermore, our work serves as a strong foundation for understanding the structural design and evolution of actin filaments and their function in motility and host cell invasion of apicomplexan parasites. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3805.map.gz emd_3805.map.gz | 6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3805-v30.xml emd-3805-v30.xml emd-3805.xml emd-3805.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3805.png emd_3805.png | 150.3 KB | ||
| Masks |  emd_3805_msk_1.map emd_3805_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-3805.cif.gz emd-3805.cif.gz | 6.6 KB | ||
| Others |  emd_3805_half_map_1.map.gz emd_3805_half_map_1.map.gz emd_3805_half_map_2.map.gz emd_3805_half_map_2.map.gz | 49.4 MB 49.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3805 http://ftp.pdbj.org/pub/emdb/structures/EMD-3805 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3805 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3805 | HTTPS FTP |
-Validation report
| Summary document |  emd_3805_validation.pdf.gz emd_3805_validation.pdf.gz | 664.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_3805_full_validation.pdf.gz emd_3805_full_validation.pdf.gz | 663.7 KB | Display | |
| Data in XML |  emd_3805_validation.xml.gz emd_3805_validation.xml.gz | 12.3 KB | Display | |
| Data in CIF |  emd_3805_validation.cif.gz emd_3805_validation.cif.gz | 14.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3805 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3805 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3805 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-3805 | HTTPS FTP |
-Related structure data
| Related structure data |  5ogwMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3805.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3805.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.14 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
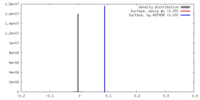
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_3805_msk_1.map emd_3805_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
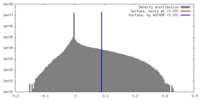
| Density Histograms |
-Half map: #1
| File | emd_3805_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
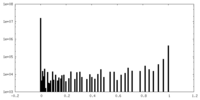
| Density Histograms |
-Half map: #2
| File | emd_3805_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
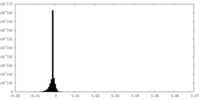
| Density Histograms |
- Sample components
Sample components
-Entire : Plasmodium falciparum actin 1 filament stabilized by jasplakinolide
| Entire | Name: Plasmodium falciparum actin 1 filament stabilized by jasplakinolide |
|---|---|
| Components |
|
-Supramolecule #1: Plasmodium falciparum actin 1 filament stabilized by jasplakinolide
| Supramolecule | Name: Plasmodium falciparum actin 1 filament stabilized by jasplakinolide type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: Filament |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Actin-1
| Macromolecule | Name: Actin-1 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 42.047676 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GAMGEEDVQA LVVDNGSGNV KAGVAGDDAP RSVFPSIVGR PKNPGIMVGM EEKDAFVGDE AQTKRGILTL KYPIEHGIVT NWDDMEKIW HHTFYNELRA APEEHPVLLT EAPLNPKGNR ERMTQIMFES FNVPAMYVAI QAVLSLYSSG RTTGIVLDSG D GVSHTVPI ...String: GAMGEEDVQA LVVDNGSGNV KAGVAGDDAP RSVFPSIVGR PKNPGIMVGM EEKDAFVGDE AQTKRGILTL KYPIEHGIVT NWDDMEKIW HHTFYNELRA APEEHPVLLT EAPLNPKGNR ERMTQIMFES FNVPAMYVAI QAVLSLYSSG RTTGIVLDSG D GVSHTVPI YEGYALPHAI MRLDLAGRDL TEYLMKILHE RGYGFSTSAE KEIVRDIKEK LCYIALNFDE EMKTSEQSSD IE KSYELPD GNIITVGNER FRCPEALFQP SFLGKEAAGI HTTTFNSIKK CDVDIRKDLY GNIVLSGGTT MYEGIGERLT RDI TTLAPS TMKIKVVAPP ERKYSVWIGG SILSSLSTFQ QMWITKEEYD ESGPSIVHRK CF UniProtKB: Actin-1 |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 5 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #3: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 3 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #4: Jasplakinolide
| Macromolecule | Name: Jasplakinolide / type: ligand / ID: 4 / Number of copies: 3 / Formula: 9UE |
|---|---|
| Molecular weight | Theoretical: 709.67 Da |
| Chemical component information |  ChemComp-9UE: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
Details: 10 mM HEPES pH 7.5, 0.2 mM CaCl2, 50 mM KCl, 4 mM MgCl2, 5 mM DTT and 0.5 mM ATP. JAS was added at a 1:1 molar ratio during polyermization. | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat-2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 97 % / Chamber temperature: 298 K / Instrument: GATAN CRYOPLUNGE 3 Details: Sample (2 uL of JAS-stabilized F-actin solution) was applied to a glow-discharged holey carbon grid, incubated for 30 s and manually blotted for 4 s from the backside with filter paper.. | ||||||||||
| Details | Twist (degree) 167.5 Rise (A) 27.4 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Details | Cs corrected microscope |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Frames/image: 1-4 / Number real images: 1634 / Average exposure time: 1.5 sec. / Average electron dose: 110.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)