[English] 日本語
 Yorodumi
Yorodumi- EMDB-36606: Cryo-EM structure of the glucagon receptor bound to beta-arrestin... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the glucagon receptor bound to beta-arrestin 1 in ligand-free state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex structure / glucagon receptor / beta-arrestin 1 / ligand-free / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein C (activated) / positive regulation of establishment of endothelial barrier / negative regulation of coagulation / regulation of glycogen metabolic process / glucagon receptor activity / negative regulation of blood coagulation / response to starvation / cellular response to glucagon stimulus / exocytosis / peptide hormone binding ...protein C (activated) / positive regulation of establishment of endothelial barrier / negative regulation of coagulation / regulation of glycogen metabolic process / glucagon receptor activity / negative regulation of blood coagulation / response to starvation / cellular response to glucagon stimulus / exocytosis / peptide hormone binding / Gamma-carboxylation of protein precursors / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / Common Pathway of Fibrin Clot Formation / Removal of aminoterminal propeptides from gamma-carboxylated proteins / Intrinsic Pathway of Fibrin Clot Formation / hormone-mediated signaling pathway / cellular response to starvation / response to nutrient / viral budding from plasma membrane / guanyl-nucleotide exchange factor activity / generation of precursor metabolites and energy / Post-translational protein phosphorylation / Cell surface interactions at the vascular wall / adenylate cyclase-activating G protein-coupled receptor signaling pathway / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / Glucagon signaling in metabolic regulation / negative regulation of inflammatory response / regulation of blood pressure / Glucagon-type ligand receptors / Golgi lumen / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / blood coagulation / glucose homeostasis / G alpha (s) signalling events / clathrin-dependent endocytosis of virus by host cell / G alpha (q) signalling events / cell surface receptor signaling pathway / endosome / host cell surface receptor binding / endoplasmic reticulum lumen / fusion of virus membrane with host plasma membrane / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / viral envelope / calcium ion binding / positive regulation of gene expression / negative regulation of apoptotic process / virion attachment to host cell / host cell plasma membrane / virion membrane / Golgi apparatus / endoplasmic reticulum / proteolysis / extracellular space / extracellular region / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Escherichia phage EcSzw-2 (virus) Escherichia phage EcSzw-2 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Chen K / Zhang C / Lin S / Zhao Q / Wu B | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Tail engagement of arrestin at the glucagon receptor. Authors: Kun Chen / Chenhui Zhang / Shuling Lin / Xinyu Yan / Heng Cai / Cuiying Yi / Limin Ma / Xiaojing Chu / Yuchen Liu / Ya Zhu / Shuo Han / Qiang Zhao / Beili Wu /  Abstract: Arrestins have pivotal roles in regulating G protein-coupled receptor (GPCR) signalling by desensitizing G protein activation and mediating receptor internalization. It has been proposed that the ...Arrestins have pivotal roles in regulating G protein-coupled receptor (GPCR) signalling by desensitizing G protein activation and mediating receptor internalization. It has been proposed that the arrestin binds to the receptor in two different conformations, 'tail' and 'core', which were suggested to govern distinct processes of receptor signalling and trafficking. However, little structural information is available for the tail engagement of the arrestins. Here we report two structures of the glucagon receptor (GCGR) bound to β-arrestin 1 (βarr1) in glucagon-bound and ligand-free states. These structures reveal a receptor tail-engaged binding mode of βarr1 with many unique features, to our knowledge, not previously observed. Helix VIII, instead of the receptor core, has a major role in accommodating βarr1 by forming extensive interactions with the central crest of βarr1. The tail-binding pose is further defined by a close proximity between the βarr1 C-edge and the receptor helical bundle, and stabilized by a phosphoinositide derivative that bridges βarr1 with helices I and VIII of GCGR. Lacking any contact with the arrestin, the receptor core is in an inactive state and loosely binds to glucagon. Further functional studies suggest that the tail conformation of GCGR-βarr governs βarr recruitment at the plasma membrane and endocytosis of GCGR, and provides a molecular basis for the receptor forming a super-complex simultaneously with G protein and βarr to promote sustained signalling within endosomes. These findings extend our knowledge about the arrestin-mediated modulation of GPCR functionalities. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36606.map.gz emd_36606.map.gz | 59.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36606-v30.xml emd-36606-v30.xml emd-36606.xml emd-36606.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36606.png emd_36606.png | 47.7 KB | ||
| Others |  emd_36606_half_map_1.map.gz emd_36606_half_map_1.map.gz emd_36606_half_map_2.map.gz emd_36606_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36606 http://ftp.pdbj.org/pub/emdb/structures/EMD-36606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36606 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36606 | HTTPS FTP |
-Validation report
| Summary document |  emd_36606_validation.pdf.gz emd_36606_validation.pdf.gz | 842.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36606_full_validation.pdf.gz emd_36606_full_validation.pdf.gz | 842 KB | Display | |
| Data in XML |  emd_36606_validation.xml.gz emd_36606_validation.xml.gz | 12.3 KB | Display | |
| Data in CIF |  emd_36606_validation.cif.gz emd_36606_validation.cif.gz | 14.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36606 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36606 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36606 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36606 | HTTPS FTP |
-Related structure data
| Related structure data |  8jruMC  8jrvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_36606.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36606.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.071 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36606_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
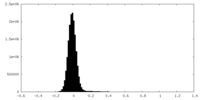
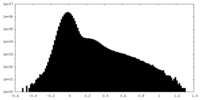
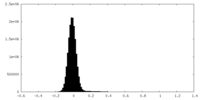
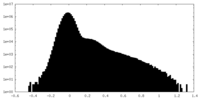
| Density Histograms |
-Half map: #1
| File | emd_36606_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : The glucagon receptor bound to beta-arrestin 1
| Entire | Name: The glucagon receptor bound to beta-arrestin 1 |
|---|---|
| Components |
|
-Supramolecule #1: The glucagon receptor bound to beta-arrestin 1
| Supramolecule | Name: The glucagon receptor bound to beta-arrestin 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: HA signal peptide,HPC4 purification tag,Glucagon receptor,C-termi...
| Macromolecule | Name: HA signal peptide,HPC4 purification tag,Glucagon receptor,C-terminal tail of Vasopressin V2 receptor type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.307141 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSMKTIIALS YIFCLVFAGA PEDQVDPRLI DGKGSGSAGS AGSQVMDFLF EKWKLYGDQC HHNLSLLPPP TELVCNRTFD KYSCWPDTP ANTTANISCP WYLPWHHKVQ HRFVFKRCGP DGQWVRGPRG QPWRDASQCQ MDGEEIEVQK EVAKMYSSFQ V MYTVGYSL ...String: GSMKTIIALS YIFCLVFAGA PEDQVDPRLI DGKGSGSAGS AGSQVMDFLF EKWKLYGDQC HHNLSLLPPP TELVCNRTFD KYSCWPDTP ANTTANISCP WYLPWHHKVQ HRFVFKRCGP DGQWVRGPRG QPWRDASQCQ MDGEEIEVQK EVAKMYSSFQ V MYTVGYSL SLGALLLALA ILGGLSKLHC TRNAIHANLF ASFVLKASSV LVIDGLLRTR YSQKIGDDLS VSTWLSDGAV AG CRVAAVF MQYGIVANYC WLLVEGLYLH NLLGLATLPE RSFFSLYLGI GWGAPMLFVV PWAVVKCLFE NVQCWTSNDN MGF WWILRF PVFLAILINF FIFVRIVQLL VAKLRARQMH HTDYKFRLAK STLTLIPLLG VHEVVFAFVT DEHAQGTLRS AKLF FDLFL SSFQGLLVAV LYCFLNKEVQ SELRRRWHRW RLGKVLWEER NTSNARGRTP PSLGPQDE(SEP)C T(TPO)A (SEP)(SEP)(SEP)LAK DTSS UniProtKB: Hemagglutinin, Vitamin K-dependent protein C, Glucagon receptor, UNIPROTKB: P30518 |
-Macromolecule #2: Beta-arrestin 1 and single-chain fragment variable 30 (scFv30)
| Macromolecule | Name: Beta-arrestin 1 and single-chain fragment variable 30 (scFv30) type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 69.173891 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGDKGTRVFK KASPNGKLTV YLGKRDFVDH IDLVEPVDGV VLVDPEYLKE RRVYVTLTAA FRYGREDLDV LGLTFRKDLF VANVQSFPP APEDKKPLTR LQERLIKKLG EHAYPFTFEI PPNLPSSVTL QPGPEDTGKA IGVDYEVKAF VAENLEEKIH K RNSVRLVI ...String: MGDKGTRVFK KASPNGKLTV YLGKRDFVDH IDLVEPVDGV VLVDPEYLKE RRVYVTLTAA FRYGREDLDV LGLTFRKDLF VANVQSFPP APEDKKPLTR LQERLIKKLG EHAYPFTFEI PPNLPSSVTL QPGPEDTGKA IGVDYEVKAF VAENLEEKIH K RNSVRLVI EKVQYAPERP GPQPTAETTR QFLMSDKPLH LEASLDKEIY YHGEPISVNV HVTNNTNKTV KKIKISVRQY AD IVLFNTA QYKVPVAMEE ADDTVAPSST FSKVYTLTPF LANNREKRGL ALDGKLKHED TNLASSTLLR EGANREILGI IVS YKVKVK LVVSRGGLLG DLASSDVAVE LPFTLMHPKP KEEPPHREVP EHETPVDTNL SDIQMTQSPS SLSASVGDRV TITC RASQS VSSAVAWYQQ KPGKAPKLLI YSASSLYSGV PSRFSGSRSG TDFTLTISSL QPEDFATYYC QQYKYVPVTF GQGTK VEIK GTTAASGSSG GSSSGAEVQL VESGGGLVQP GGSLRLSCAA SGFNVYSSSI HWVRQAPGKG LEWVASISSY YGYTYY ADS VKGRFTISAD TSKNTAYLQM NSLRAEDTAV YYCARSRQFW YSGLDYWGQG TLVTVSSAHH HHHH |
-Macromolecule #3: Nanobody 32
| Macromolecule | Name: Nanobody 32 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage EcSzw-2 (virus) Escherichia phage EcSzw-2 (virus) |
| Molecular weight | Theoretical: 13.867408 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAQVQLQESG GGLVQAGGSL RLSCVVSGFF FDTVTMAWYR RAPGKHRELV ASATAGGTTT YADSVKDRFT ISRDNAKNTV YLQMNSLKP EDTAVYYCNT FVRSLSWGQG TQVTVSSHHH HHHEPEA |
-Macromolecule #4: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(o...
| Macromolecule | Name: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate type: ligand / ID: 4 / Number of copies: 1 / Formula: PIO |
|---|---|
| Molecular weight | Theoretical: 746.566 Da |
| Chemical component information |  ChemComp-PIO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 250907 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)