+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Portal-tail complex of phage GP4 | |||||||||||||||
 Map data Map data | portal tail complex of phage GP4 | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Complex / VIRAL PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||||||||
| Biological species |  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||||||||
 Authors Authors | Liu H / Chen W | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
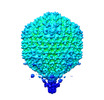
 Citation Citation |  Journal: J Mol Biol / Year: 2023 Journal: J Mol Biol / Year: 2023Title: Asymmetric Structure of Podophage GP4 Reveals a Novel Architecture of Three Types of Tail Fibers. Authors: Jing Zheng / Wenyuan Chen / Hao Xiao / Fan Yang / Jingdong Song / Lingpeng Cheng / Hongrong Liu /  Abstract: Bacteriophage tail fibers (or called tail spikes) play a critical role in the early stage of infection by binding to the bacterial surface. Podophages with known structures usually possess one or two ...Bacteriophage tail fibers (or called tail spikes) play a critical role in the early stage of infection by binding to the bacterial surface. Podophages with known structures usually possess one or two types of fibers. Here, we resolved an asymmetric structure of the podophage GP4 to near-atomic resolution by cryo-EM. Our structure revealed a symmetry-mismatch relationship between the components of the GP4 tail with previously unseen topologies. In detail, two dodecameric adaptors (adaptors I and II), a hexameric nozzle, and a tail needle form a conserved tail body connected to a dodecameric portal occupying a unique vertex of the icosahedral head. However, five chain-like extended fibers (fiber I) and five tulip-like short fibers (fiber II) are anchored to a 15-fold symmetric fiber-tail adaptor, encircling the adaptor I, and six bamboo-like trimeric fibers (fiber III) are connected to the nozzle. Five fibers I, each composed of five dimers of the protein gp80 linked by an elongated rope protein, are attached to the five edges of the tail vertex of the icosahedral head. In this study, we identified a new structure of the podophage with three types of tail fibers, and such phages with different types of fibers may have a broad host range and/or infect host cells with considerably high efficiency, providing evolutionary advantages in harsh environments. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36463.map.gz emd_36463.map.gz | 153.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36463-v30.xml emd-36463-v30.xml emd-36463.xml emd-36463.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36463.png emd_36463.png | 101.7 KB | ||
| Filedesc metadata |  emd-36463.cif.gz emd-36463.cif.gz | 6.5 KB | ||
| Others |  emd_36463_half_map_1.map.gz emd_36463_half_map_1.map.gz emd_36463_half_map_2.map.gz emd_36463_half_map_2.map.gz | 221.9 MB 221.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36463 http://ftp.pdbj.org/pub/emdb/structures/EMD-36463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36463 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36463 | HTTPS FTP |
-Validation report
| Summary document |  emd_36463_validation.pdf.gz emd_36463_validation.pdf.gz | 985 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36463_full_validation.pdf.gz emd_36463_full_validation.pdf.gz | 984.6 KB | Display | |
| Data in XML |  emd_36463_validation.xml.gz emd_36463_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_36463_validation.cif.gz emd_36463_validation.cif.gz | 19 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36463 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36463 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36463 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36463 | HTTPS FTP |
-Related structure data
| Related structure data |  8jovMC  8jouC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36463.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36463.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | portal tail complex of phage GP4 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.27 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map1
| File | emd_36463_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map2
| File | emd_36463_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ralstonia phage GP4
| Entire | Name:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Ralstonia phage GP4
| Supramolecule | Name: Ralstonia phage GP4 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 2282904 / Sci species name: Ralstonia phage GP4 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Portal protein
| Macromolecule | Name: Portal protein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
| Molecular weight | Theoretical: 87.929188 KDa |
| Sequence | String: MFDLRDDQNT RVIADHPEAR MPTEEPGKDD APPSHPLDSE EMKELHRRLR SYLHQELDRQ AENRFQMAVD EDYYDSIQWT EADAQVLKE RGQAPLVFNV IAQSVNWIIG SEKRGRTDFN ILPRKKADAK PAEAKTKLLK YLSDVNRLPF HRSRSFEDTV K VGLGWIED ...String: MFDLRDDQNT RVIADHPEAR MPTEEPGKDD APPSHPLDSE EMKELHRRLR SYLHQELDRQ AENRFQMAVD EDYYDSIQWT EADAQVLKE RGQAPLVFNV IAQSVNWIIG SEKRGRTDFN ILPRKKADAK PAEAKTKLLK YLSDVNRLPF HRSRSFEDTV K VGLGWIED SYDDSTDGEP IYSRYESWRN VIFDSASTEL DGTDMRYIFR PKWLDVDVAC ALVPDRADEI KKAAVAAERY GN YSEEDGD EAMDWAEFDR DTYSQSRTVS THKRQRVRLI ECWYRKPMRA TKFLSGDLRG ETLDESNPAH VEARDSGLYS LGE RITMRV RVAIFTSRDL LFEGASPFRH NRFSLTPIWG FRRGRDNLPY GVIRWMRDIQ DDINKRASKA LYILSSNKVV MDEG AVEDI EEFREEVARP DAVLVKKPGK QIELNVDREL AAAHMDMMSR DIQMLQQVGG VTDELMGRST NAVSGVAIQA RQEQG TVAT NKLFDNLRFA VQMQGEIQLS LIEQFVTEEK TFRITNERGK ADFITVNDGL PENDIVRTKA DFIIGESDWR ATYRQA ASE QLSQMIMKMP PQVGLVMLDL WADSTDLPNR DEIVKRIRQI NGMRDPDATE PTPEELQQQQ AAAEQAQAQK AMFMAEL SE KQGKADKAQA DAVAAQANAD LRAAQAEQVR RQTVNTNVTS IAAAMEAATA IVTMPTIAKV GDAVLVEAGY ENNGIAPA G GLHTPAPQQA VNPAAQGLPP QQPQPQQPQE PAVSPSPEQL NGAAPSNTGE PVTPDQTLQQ UniProtKB: Portal protein |
-Macromolecule #2: Putative tail fiber protein
| Macromolecule | Name: Putative tail fiber protein / type: protein_or_peptide / ID: 2 / Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
| Molecular weight | Theoretical: 45.894781 KDa |
| Sequence | String: MAAVQFANNA ASRLVGPLAP SGTSLTVTPG DGALFPTLGA GDWFMATLIR SDGAREIVKV TSRTVDAMSI TRAQEGSSAL SFNPNDRIE ARLTAGELGD FRDGIASAQS AASAAQGTAD GKISKAGDTM TGNLNMEGAA PIITFRETDQ AAPAGRWRLV A DGGNWSLR ...String: MAAVQFANNA ASRLVGPLAP SGTSLTVTPG DGALFPTLGA GDWFMATLIR SDGAREIVKV TSRTVDAMSI TRAQEGSSAL SFNPNDRIE ARLTAGELGD FRDGIASAQS AASAAQGTAD GKISKAGDTM TGNLNMEGAA PIITFRETDQ AAPAGRWRLV A DGGNWSLR RSTAGEFASE NSTMWFGPDD SVHFLNDILI GRGNLGSVYD ALASKATSAA LASGLSAKPD ADGVSVTGFV NG NFYEPYF RKSSDNTVRR LVANTSANGI ALSWSGSFLS RTIDNTATAT IWDTANAPGA GTTNLDKTFY CGDGTRIGRS WVP GSGFLS ITVDGTNYGI SISASDERLK REIAPSEASA LSKLGRIELF SFRYKEGNAF LDPSQHHDIG FIAQQLASVD PTFV AGGGE TMLSPNLQPI VATLVKAVQE LRSQVDALKA QVGA UniProtKB: Putative tail fiber protein |
-Macromolecule #3: Virion associated protein
| Macromolecule | Name: Virion associated protein / type: protein_or_peptide / ID: 3 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
| Molecular weight | Theoretical: 23.545262 KDa |
| Sequence | String: MAFPASVVLS RAATLLQDED HERWTVDELL EWLTDGTREI VVRKPSAYMK TTTAALVAGS KQALPEDAIQ LIDVPRNLKT DGSPGRAVT ATDRRLLDTE NPDWHSMKPA GQIRHYTYDS NVPTVFYTYP PAAAGVQVEL VCAWRHPALT TQNDVVQMGA E FVSALVSW ...String: MAFPASVVLS RAATLLQDED HERWTVDELL EWLTDGTREI VVRKPSAYMK TTTAALVAGS KQALPEDAIQ LIDVPRNLKT DGSPGRAVT ATDRRLLDTE NPDWHSMKPA GQIRHYTYDS NVPTVFYTYP PAAAGVQVEL VCAWRHPALT TQNDVVQMGA E FVSALVSW CLYRASSKDS EFANGAVAAA HYSAFSDALG AQATGTPTTQ AAAAAAAAGA AQ UniProtKB: Virion associated protein |
-Macromolecule #4: Virion-associated phage protein
| Macromolecule | Name: Virion-associated phage protein / type: protein_or_peptide / ID: 4 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
| Molecular weight | Theoretical: 62.764598 KDa |
| Sequence | String: MTTLKLTAYS GEVPRTLPRL LPDTASQRAL NVRLDNGGLT PTRQPRFEAN ISVDNAKTIY KHNGAWLAWQ NVVHAAPGPV AQDRLYYMG DGKPKMIVDG TTYDLAVPMP TAAPALTVTG TGTGNVTSIA YVYTFVTAFG EESEPSALSN VAGWQSGQTR T LTGIQAPP ...String: MTTLKLTAYS GEVPRTLPRL LPDTASQRAL NVRLDNGGLT PTRQPRFEAN ISVDNAKTIY KHNGAWLAWQ NVVHAAPGPV AQDRLYYMG DGKPKMIVDG TTYDLAVPMP TAAPALTVTG TGTGNVTSIA YVYTFVTAFG EESEPSALSN VAGWQSGQTR T LTGIQAPP AGRNITKQRF YRSQTGSGGT DLFFIEERAA SAANFVDTHA TNDFGEMLPS LEYNAPPDGL KGLISLPNGM MA AFTGKDL YFCEPFIPHA WPEKYILTMD YQIVALGAYG TTIVVMTEGL PYIVSGTAPE NMQQQRVELN LPCINARGVI DLG YSVAYP SHDGLVMAGS NGMQVITEQL MTRNDWMKTG PGNIVGGQFN GRYFASYEYI EPSGAAFSGT LIFDTTGAAP FIIR SNHKA DAFFHELQTG ALYFLVGKEI FEWDALGQVN ETLSWRSKQF VLPMPTNFGA ILIEGSTAAS EEEQAAYDAE RQRIE VENA TNFALPSIGG EMNGAEVNLF AVNGDMMQRL PAEGFVSVSI YADGKLVKTV SKMNRMARLP SGFLARIWEI EVNSNI NIS DIVLATTGQE LRNV UniProtKB: Virion-associated phage protein |
-Macromolecule #5: gp81 of phage GP4
| Macromolecule | Name: gp81 of phage GP4 / type: protein_or_peptide / ID: 5 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Ralstonia phage GP4 (virus) Ralstonia phage GP4 (virus) |
| Molecular weight | Theoretical: 23.197357 KDa |
| Sequence | String: MKPLDVFMPI IHRFAPGCPE PTAFAAIREA AIKFCERTRL WRCDDEFNVG ADECAEVAVP YGAALYEMEL VQFNGRNMRP VSTQWLDEQ VPDWRTTTQS GQAQYVTQQS EDTLTFVPAE AGSVRVYGLL KPTLDADSLP DLLADTYRKT IADGALSELL I IPGKAWMS ...String: MKPLDVFMPI IHRFAPGCPE PTAFAAIREA AIKFCERTRL WRCDDEFNVG ADECAEVAVP YGAALYEMEL VQFNGRNMRP VSTQWLDEQ VPDWRTTTQS GQAQYVTQQS EDTLTFVPAE AGSVRVYGLL KPTLDADSLP DLLADTYRKT IADGALSELL I IPGKAWMS ADLAVFFGTR FDRELDRLST KTIKGQQRAP VRTRAQFF UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.8000000000000003 µm / Nominal defocus min: 0.1 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 39883 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X