[English] 日本語
 Yorodumi
Yorodumi- EMDB-3636: multi-drug efflux; membrane transport; RND superfamily; Drug resi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3636 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | multi-drug efflux; membrane transport; RND superfamily; Drug resistance | |||||||||
 Map data Map data | AcrABZ-TolC pump with inhibitor MBX3132 bound AcrB in the TTT state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | multi-drug efflux / membrane transport / RND superfamily / Drug resistance / membrane protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationMacAB-TolC complex / alkane transmembrane transporter activity / alkane transport / enterobactin transport / enterobactin transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / periplasmic side of plasma membrane / efflux pump complex / Iron assimilation using enterobactin / bile acid transmembrane transporter activity ...MacAB-TolC complex / alkane transmembrane transporter activity / alkane transport / enterobactin transport / enterobactin transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / periplasmic side of plasma membrane / efflux pump complex / Iron assimilation using enterobactin / bile acid transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the plasma membrane / Antimicrobial resistance / bile acid and bile salt transport / porin activity / Secretion of toxins / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / monoatomic ion channel activity / fatty acid transport / xenobiotic transport / cell outer membrane / response to toxic substance / outer membrane-bounded periplasmic space / monoatomic ion transmembrane transport / response to xenobiotic stimulus / response to antibiotic / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
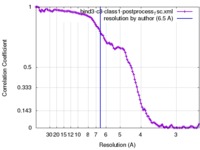
| Method | single particle reconstruction / cryo EM / Resolution: 6.5 Å | |||||||||
 Authors Authors | Wang Z / Fan G | |||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: An allosteric transport mechanism for the AcrAB-TolC multidrug efflux pump. Authors: Zhao Wang / Guizhen Fan / Corey F Hryc / James N Blaza / Irina I Serysheva / Michael F Schmid / Wah Chiu / Ben F Luisi / Dijun Du /   Abstract: Bacterial efflux pumps confer multidrug resistance by transporting diverse antibiotics from the cell. In Gram-negative bacteria, some of these pumps form multi-protein assemblies that span the cell ...Bacterial efflux pumps confer multidrug resistance by transporting diverse antibiotics from the cell. In Gram-negative bacteria, some of these pumps form multi-protein assemblies that span the cell envelope. Here, we report the near-atomic resolution cryoEM structures of the AcrAB-TolC multidrug efflux pump in resting and drug transport states, revealing a quaternary structural switch that allosterically couples and synchronizes initial ligand binding with channel opening. Within the transport-activated state, the channel remains open even though the pump cycles through three distinct conformations. Collectively, our data provide a dynamic mechanism for the assembly and operation of the AcrAB-TolC pump. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3636.map.gz emd_3636.map.gz | 105.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3636-v30.xml emd-3636-v30.xml emd-3636.xml emd-3636.xml | 20.5 KB 20.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3636_fsc.xml emd_3636_fsc.xml | 10.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_3636.png emd_3636.png | 78.1 KB | ||
| Filedesc metadata |  emd-3636.cif.gz emd-3636.cif.gz | 8.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3636 http://ftp.pdbj.org/pub/emdb/structures/EMD-3636 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3636 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3636 | HTTPS FTP |
-Related structure data
| Related structure data |  5ng5MC  8636C  8640C  5nc5C  5o66C  5v5sC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3636.map.gz / Format: CCP4 / Size: 113.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3636.map.gz / Format: CCP4 / Size: 113.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | AcrABZ-TolC pump with inhibitor MBX3132 bound AcrB in the TTT state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.29 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : AcrABTolC in apo state
| Entire | Name: AcrABTolC in apo state |
|---|---|
| Components |
|
-Supramolecule #1: AcrABTolC in apo state
| Supramolecule | Name: AcrABTolC in apo state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Multidrug efflux pump subunit AcrA
| Macromolecule | Name: Multidrug efflux pump subunit AcrA / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 39.80066 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: CDDKQAQQGG QQMPAVGVVT VKTEPLQITT ELPGRTSAYR IAEVRPQVSG IILKRNFKEG SDIEAGVSLY QIDPATYQAT YDSAKGDLA KAQAAANIAQ LTVNRYQKLL GTQYISKQEY DQALADAQQA NAAVTAAKAA VETARINLAY TKVTSPISGR I GKSNVTEG ...String: CDDKQAQQGG QQMPAVGVVT VKTEPLQITT ELPGRTSAYR IAEVRPQVSG IILKRNFKEG SDIEAGVSLY QIDPATYQAT YDSAKGDLA KAQAAANIAQ LTVNRYQKLL GTQYISKQEY DQALADAQQA NAAVTAAKAA VETARINLAY TKVTSPISGR I GKSNVTEG ALVQNGQATA LATVQQLDPI YVDVTQSSND MMRLKQELAN GTLKQENGKA KVSLITSDGI KFPQDGTLEF SD VTVDQTT GSITLRAIFP NPDHTMMPGM FVRARLEEGL NPNAILVPQQ GVTRTPRGDA TVLVVGADDK VETRPIVASQ AIG DKWLVT EGLKAGDRVV ISGLQKVRPG VQVKAQEVTA DNNQQAASGA QPEQSKS UniProtKB: Multidrug efflux pump subunit AcrA |
-Macromolecule #2: Outer membrane protein TolC
| Macromolecule | Name: Outer membrane protein TolC / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.783355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKLLPILIG LSLSGFSSLS QAENLMQVYQ QARLSNPELR KSAADRDAAF EKINEARSPL LPQLGLGADY TYSNGYRDAN GINSNATSA SLQLTQSIFD MSKWRALTLQ EKAAGIQDVT YQTDQQTLIL NTATAYFNVL NAIDVLSYTQ AQKEAIYRQL D QTTQRFNV ...String: MKKLLPILIG LSLSGFSSLS QAENLMQVYQ QARLSNPELR KSAADRDAAF EKINEARSPL LPQLGLGADY TYSNGYRDAN GINSNATSA SLQLTQSIFD MSKWRALTLQ EKAAGIQDVT YQTDQQTLIL NTATAYFNVL NAIDVLSYTQ AQKEAIYRQL D QTTQRFNV GLVAITDVQN ARAQYDTVLA NEVTARNNLD NAVEQLRQIT GNYYPELAAL NVENFKTDKP QPVNALLKEA EK RNLSLLQ ARLSQDLARE QIRQAQDGHL PTLDLTASTG ISDTSYSGSK TRGAAGTQYD DSNMGQNKVG LSFSLPIYQG GMV NSQVKQ AQYNFVGASE QLESAHRSVV QTVRSSFNNI NASISSINAY KQAVVSAQSS LDAMEAGYSV GTRTIVDVLD ATTT LYNAK QELANARYNY LINQLNIKSA LGTLNEQDLL ALNNALSKPV STNPENVAPQ TPEQNAIADG YAPDSPAPVV QQTSA RTTT SNGHNPFRN UniProtKB: Outer membrane protein TolC |
-Macromolecule #3: Multidrug efflux pump subunit AcrB
| Macromolecule | Name: Multidrug efflux pump subunit AcrB / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 113.66518 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR ...String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR TSGVGDVQLF GSQYAMRIWM NPNELNKFQL TPVDVITAIK AQNAQVAAGQ LGGTPPVKGQ QLNASIIAQT RL TSTEEFG KILLKVNQDG SRVLLRDVAK IELGGENYDI IAEFNGQPAS GLGIKLATGA NALDTAAAIR AELAKMEPFF PSG LKIVYP YDTTPFVKIS IHEVVKTLVE AIILVFLVMY LFLQNFRATL IPTIAVPVVL LGTFAVLAAF GFSINTLTMF GMVL AIGLL VDDAIVVVEN VERVMAEEGL PPKEATRKSM GQIQGALVGI AMVLSAVFVP MAFFGGSTGA IYRQFSITIV SAMAL SVLV ALILTPALCA TMLKPIAKGD HGEGKKGFFG WFNRMFEKST HHYTDSVGGI LRSTGRYLVL YLIIVVGMAY LFVRLP SSF LPDEDQGVFM TMVQLPAGAT QERTQKVLNE VTHYYLTKEK NNVESVFAVN GFGFAGRGQN TGIAFVSLKD WADRPGE EN KVEAITMRAT RAFSQIKDAM VFAFNLPAIV ELGTATGFDF ELIDQAGLGH EKLTQARNQL LAEAAKHPDM LTSVRPNG L EDTPQFKIDI DQEKAQALGV SINDINTTLG AAWGGSYVND FIDRGRVKKV YVMSEAKYRM LPDDIGDWYV RAADGQMVP FSAFSSSRWE YGSPRLERYN GLPSMEILGQ AAPGKSTGEA MELMEQLASK LPTGVGYDWT GMSYQERLSG NQAPSLYAIS LIVVFLCLA ALYESWSIPF SVMLVVPLGV IGALLAATFR GLTNDVYFQV GLLTTIGLSA KNAILIVEFA KDLMDKEGKG L IEATLDAV RMRLRPILMT SLAFILGVMP LVISTGAGSG AQNAVGTGVM GGMVTATVLA IFFVPVFFVV VRRRFSRKNE DI EHSHTVD HH UniProtKB: Multidrug efflux pump subunit AcrB |
-Macromolecule #4: Multidrug efflux pump accessory protein AcrZ
| Macromolecule | Name: Multidrug efflux pump accessory protein AcrZ / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5.995157 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLELLKSLVF AVIMVPVVMA IILGLIYGLG EVFNIFSGVG KKDQPGQNHH HHHH UniProtKB: Multidrug efflux pump accessory protein AcrZ |
-Macromolecule #5: 6-[2-(3,4-dimethoxyphenyl)ethylsulfanyl]-8-[4-(2-methoxyethyl)pip...
| Macromolecule | Name: 6-[2-(3,4-dimethoxyphenyl)ethylsulfanyl]-8-[4-(2-methoxyethyl)piperazin-1-yl]-3,3-dimethyl-1,4-dihydropyrano[3,4-c]pyridine-5-carbonitrile type: ligand / ID: 5 / Number of copies: 3 / Formula: 5QF |
|---|---|
| Molecular weight | Theoretical: 526.691 Da |
| Chemical component information | 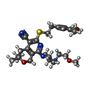 ChemComp-5QF: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 20 K / Instrument: FEI VITROBOT MARK IV Details: a 3ul aliquot at a concentration of 2 mg per ml was applied onto glow-discharged holey carbon grid (Quantifoil Au R1.21.3, 300 mesh). |
| Details | AcrABTolC complex in apo state |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Crystal structures were rigid-body fit into the density map and model optimization was then carried out with Phenix real-space refine. Due to the weaker resolution, stronger stereochemical and secondary structure restraints were used to ensure that alpha-helices and beta-sheets did not deviate far from their expected geometry. Manual adjustments were kept to a minimum to reduce human bias in the modeling procedure, with Coot only being used to fix obvious errors such as C-beta deviations. A final check of MolProbity and cross correlation was done to ensure model quality. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-5ng5: |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)