+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Mycolicibacterium smegmatis F1-ATPase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / F-ATP synthase / cryo-EM / mycobacteria / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationproton motive force-driven plasma membrane ATP synthesis / proton-transporting ATP synthase complex, catalytic core F(1) / H+-transporting two-sector ATPase / proton-transporting ATPase activity, rotational mechanism / proton-transporting ATP synthase activity, rotational mechanism / ADP binding / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) | |||||||||
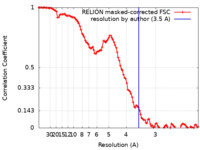
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Wong CF / Saw W-G / Grueber G | |||||||||
| Funding support |  Singapore, 1 items Singapore, 1 items
| |||||||||
 Citation Citation |  Journal: Antimicrob Agents Chemother / Year: 2022 Journal: Antimicrob Agents Chemother / Year: 2022Title: Structural Elements Involved in ATP Hydrolysis Inhibition and ATP Synthesis of Tuberculosis and Nontuberculous Mycobacterial F-ATP Synthase Decipher New Targets for Inhibitors. Authors: Chui Fann Wong / Wuan-Geok Saw / Sandip Basak / Mio Sano / Hiroshi Ueno / Hwee Wen Kerk / Dennis Litty / Priya Ragunathan / Thomas Dick / Volker Müller / Hiroyuki Noji / Gerhard Grüber /     Abstract: The FF-ATP synthase is required for the viability of tuberculosis (TB) and nontuberculous mycobacteria (NTM) and has been validated as a drug target. Here, we present the cryo-EM structures of the ...The FF-ATP synthase is required for the viability of tuberculosis (TB) and nontuberculous mycobacteria (NTM) and has been validated as a drug target. Here, we present the cryo-EM structures of the Mycobacterium smegmatis F-ATPase and the FF-ATP synthase with different nucleotide occupation within the catalytic sites and visualize critical elements for latent ATP hydrolysis and efficient ATP synthesis. Mutational studies reveal that the extended C-terminal domain (αCTD) of subunit α is the main element for the self-inhibition mechanism of ATP hydrolysis for TB and NTM bacteria. Rotational studies indicate that the transition between the inhibition state by the αCTD and the active state is a rapid process. We demonstrate that the unique mycobacterial γ-loop and subunit δ are critical elements required for ATP formation. The data underline that these mycobacterium-specific elements of α, γ, and δ are attractive targets, providing a platform for the discovery of species-specific inhibitors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33614.map.gz emd_33614.map.gz | 7.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33614-v30.xml emd-33614-v30.xml emd-33614.xml emd-33614.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33614_fsc.xml emd_33614_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_33614.png emd_33614.png | 118.6 KB | ||
| Masks |  emd_33614_msk_1.map emd_33614_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-33614.cif.gz emd-33614.cif.gz | 7 KB | ||
| Others |  emd_33614_half_map_1.map.gz emd_33614_half_map_1.map.gz emd_33614_half_map_2.map.gz emd_33614_half_map_2.map.gz | 81 MB 80.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33614 http://ftp.pdbj.org/pub/emdb/structures/EMD-33614 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33614 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33614 | HTTPS FTP |
-Validation report
| Summary document |  emd_33614_validation.pdf.gz emd_33614_validation.pdf.gz | 909.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_33614_full_validation.pdf.gz emd_33614_full_validation.pdf.gz | 909 KB | Display | |
| Data in XML |  emd_33614_validation.xml.gz emd_33614_validation.xml.gz | 17.2 KB | Display | |
| Data in CIF |  emd_33614_validation.cif.gz emd_33614_validation.cif.gz | 22.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33614 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33614 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33614 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-33614 | HTTPS FTP |
-Related structure data
| Related structure data |  7y5aMC  7y5bC  7y5cC  7y5dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33614.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33614.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_33614_msk_1.map emd_33614_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33614_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33614_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : F1-ATPase
| Entire | Name: F1-ATPase |
|---|---|
| Components |
|
-Supramolecule #1: F1-ATPase
| Supramolecule | Name: F1-ATPase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: mc(2)155 |
| Molecular weight | Theoretical: 380 KDa |
-Macromolecule #1: ATP synthase subunit alpha
| Macromolecule | Name: ATP synthase subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: H+-transporting two-sector ATPase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 58.951461 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) |
| Sequence | String: MAELTISAAD IEGAIEDYVS SFSADTEREE IGTVIDAGDG IAHVEGLPSV MTQELLEFPG GVLGVALNLD EHSVGAVILG EFEKIEEGQ QVKRTGEVLS VPVGDAFLGR VVNPLGQPID GQGDIAAETR RALELQAPSV VQRQSVSEPL QTGIKAIDAM T PIGRGQRQ ...String: MAELTISAAD IEGAIEDYVS SFSADTEREE IGTVIDAGDG IAHVEGLPSV MTQELLEFPG GVLGVALNLD EHSVGAVILG EFEKIEEGQ QVKRTGEVLS VPVGDAFLGR VVNPLGQPID GQGDIAAETR RALELQAPSV VQRQSVSEPL QTGIKAIDAM T PIGRGQRQ LIIGDRKTGK TAVCVDTILN QREAWLTGDP KQQVRCVYVA IGQKGTTIAS VKRALEEGGA MEYTTIVAAP AS DAAGFKW LAPYTGSAIG QHWMYNGKHV LIVFDDLSKQ ADAYRAISLL LRRPPGREAF PGDVFYLHSR LLERCAKLSD ELG GGSMTG LPIIETKAND ISAFIPTNVI SITDGQCFLE SDLFNQGVRP AINVGVSVSR VGGAAQIKAM KEVAGSLRLD LSQY RELEA FAAFASDLDA ASKAQLDRGA RLVELLKQPQ YSPLAVEEQV VAIFLGTQGH LDSVPVEDVQ RFESELLEHV KASHS DIFD GIRETKKLSE EAEEKLVSVI NEFKKGFQAS DGSSVVVSEN AEALDPEDLE KESVKVRKPA PKKA UniProtKB: ATP synthase subunit alpha |
-Macromolecule #2: ATP synthase subunit beta
| Macromolecule | Name: ATP synthase subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: H+-transporting two-sector ATPase |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 52.499332 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) |
| Sequence | String: MHHHHHHTAT AEKTAGRVVR ITGPVVDVEF PRGSVPELFN ALHAEITFGA LAKTLTLEVA QHLGDSLVRC ISMQPTDGLV RGVEVTDTG ASISVPVGDG VKGHVFNALG DCLDDPGYGK DFEHWSIHRK PPAFSDLEPR TEMLETGLKV VDLLTPYVRG G KIALFGGA ...String: MHHHHHHTAT AEKTAGRVVR ITGPVVDVEF PRGSVPELFN ALHAEITFGA LAKTLTLEVA QHLGDSLVRC ISMQPTDGLV RGVEVTDTG ASISVPVGDG VKGHVFNALG DCLDDPGYGK DFEHWSIHRK PPAFSDLEPR TEMLETGLKV VDLLTPYVRG G KIALFGGA GVGKTVLIQE MINRIARNFG GTSVFAGVGE RTREGNDLWV ELADANVLKD TALVFGQMDE PPGTRMRVAL SA LTMAEFF RDEQGQDVLL FIDNIFRFTQ AGSEVSTLLG RMPSAVGYQP TLADEMGELQ ERITSTRGRS ITSMQAVYVP ADD YTDPAP ATTFAHLDAT TELSRAVFSK GIFPAVDPLA SSSTILDPAI VGDEHYRVAQ EVIRILQRYK DLQDIIAILG IDEL SEEDK QLVNRARRIE RFLSQNMMAA EQFTGQPGST VPLKETIEAF DKLTKGEFDH LPEQAFFLIG GLDDLAKKAE SLGAK L UniProtKB: ATP synthase subunit beta |
-Macromolecule #3: ATP synthase gamma chain
| Macromolecule | Name: ATP synthase gamma chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 Mycolicibacterium smegmatis (bacteria) / Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 33.439836 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) |
| Sequence | String: MAATLRELRG RIRSAGSIKK ITKAQELIAT SRIAKAQARV EAARPYAAEI TNMLTELAGA SALDHPLLVE RKQPKRAGVL VVSSDRGLC GAYNANVLRR AEELFSLLRD EGKDPVLYVV GRKALGYFSF RQRTVVESWT GFSERPTYEN AREIADTLVN A FMAGADDE ...String: MAATLRELRG RIRSAGSIKK ITKAQELIAT SRIAKAQARV EAARPYAAEI TNMLTELAGA SALDHPLLVE RKQPKRAGVL VVSSDRGLC GAYNANVLRR AEELFSLLRD EGKDPVLYVV GRKALGYFSF RQRTVVESWT GFSERPTYEN AREIADTLVN A FMAGADDE GDDAGADGIL GVDELHIVFT EFRSMLSQTA VARRAAPMEV EYVGEVETGP RTLYSFEPDP ETLFDALLPR YI ATRVYAA LLEAAASESA SRRRAMKSAT DNADDLIKAL TLAANRERQA QITQEISEIV GGANALAGSK UniProtKB: ATP synthase gamma chain |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 3 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 5 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 2 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 1 / Number real images: 2318 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)