[English] 日本語
 Yorodumi
Yorodumi- EMDB-31278: Annealing synchronizes 70S ribosome into minimum-energy conformation -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31278 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Annealing synchronizes 70S ribosome into minimum-energy conformation | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.92 Å | |||||||||
 Authors Authors | Su X / Chu X / Shen Q | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Annealing synchronizes the 70 ribosome into a minimum-energy conformation. Authors: Xiaofeng Chu / Xin Su / Mingdong Liu / Li Li / Tianhao Li / Yicheng Qin / Guoliang Lu / Lei Qi / Yunhui Liu / Jinzhong Lin / Qing-Tao Shen /  Abstract: Researchers commonly anneal metals, alloys, and semiconductors to repair defects and improve microstructures via recrystallization. Theoretical studies indicate that simulated annealing on biological ...Researchers commonly anneal metals, alloys, and semiconductors to repair defects and improve microstructures via recrystallization. Theoretical studies indicate that simulated annealing on biological macromolecules helps predict the final structures with minimum free energy. Experimental validation of this homogenizing effect and further exploration of its applications are fascinating scientific questions that remain elusive. Here, we chose the apo-state 70 ribosome from as a model, wherein the 30 subunit undergoes a thermally driven intersubunit rotation and exhibits substantial structural flexibility as well as distinct free energy. We experimentally demonstrate that annealing at a fast cooling rate enhances the 70 ribosome homogeneity and improves local resolution on the 30 subunit. After annealing, the 70 ribosome is in a nonrotated state with respect to corresponding intermediate structures in unannealed or heated ribosomes. Manifold-based analysis further indicates that the annealed 70 ribosome takes a narrow conformational distribution and exhibits a minimum-energy state in the free-energy landscape. Our experimental results offer a facile yet robust approach to enhance protein stability, which is ideal for high-resolution cryogenic electron microscopy. Beyond structure determination, annealing shows great potential for synchronizing proteins on a single-molecule level and can be extended to study protein folding and explore conformational and energy landscapes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31278.map.gz emd_31278.map.gz | 120.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31278-v30.xml emd-31278-v30.xml emd-31278.xml emd-31278.xml | 7.6 KB 7.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31278.png emd_31278.png | 126.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31278 http://ftp.pdbj.org/pub/emdb/structures/EMD-31278 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31278 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31278 | HTTPS FTP |
-Validation report
| Summary document |  emd_31278_validation.pdf.gz emd_31278_validation.pdf.gz | 333.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_31278_full_validation.pdf.gz emd_31278_full_validation.pdf.gz | 333.1 KB | Display | |
| Data in XML |  emd_31278_validation.xml.gz emd_31278_validation.xml.gz | 7.4 KB | Display | |
| Data in CIF |  emd_31278_validation.cif.gz emd_31278_validation.cif.gz | 8.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31278 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31278 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31278 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-31278 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_31278.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31278.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
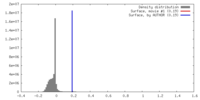
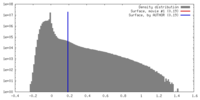
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 70S ribosome
| Entire | Name: 70S ribosome |
|---|---|
| Components |
|
-Supramolecule #1: 70S ribosome
| Supramolecule | Name: 70S ribosome / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.92 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 200000 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)