+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30445 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | human KCNQ2 in complex with retigabine | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ion channel / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationaxon initial segment / Voltage gated Potassium channels / node of Ranvier / Interaction between L1 and Ankyrins / voltage-gated monoatomic cation channel activity / ankyrin binding / action potential / voltage-gated potassium channel activity / voltage-gated potassium channel complex / potassium ion transmembrane transport ...axon initial segment / Voltage gated Potassium channels / node of Ranvier / Interaction between L1 and Ankyrins / voltage-gated monoatomic cation channel activity / ankyrin binding / action potential / voltage-gated potassium channel activity / voltage-gated potassium channel complex / potassium ion transmembrane transport / nervous system development / chemical synaptic transmission / calmodulin binding / synapse / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Li X / Lv D | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2021 Journal: Cell Res / Year: 2021Title: Molecular basis for ligand activation of the human KCNQ2 channel. Authors: Xiaoxiao Li / Qiansen Zhang / Peipei Guo / Jie Fu / Lianghe Mei / Dashuai Lv / Jiangqin Wang / Dongwu Lai / Sheng Ye / Huaiyu Yang / Jiangtao Guo /  Abstract: The voltage-gated potassium channel KCNQ2 is responsible for M-current in neurons and is an important drug target to treat epilepsy, pain and several other diseases related to neuronal hyper- ...The voltage-gated potassium channel KCNQ2 is responsible for M-current in neurons and is an important drug target to treat epilepsy, pain and several other diseases related to neuronal hyper-excitability. A list of synthetic compounds have been developed to directly activate KCNQ2, yet our knowledge of their activation mechanism is limited, due to lack of high-resolution structures. Here, we report cryo-electron microscopy (cryo-EM) structures of the human KCNQ2 determined in apo state and in complex with two activators, ztz240 or retigabine, which activate KCNQ2 through different mechanisms. The activator-bound structures, along with electrophysiology analysis, reveal that ztz240 binds at the voltage-sensing domain and directly stabilizes it at the activated state, whereas retigabine binds at the pore domain and activates the channel by an allosteric modulation. By accurately defining ligand-binding sites, these KCNQ2 structures not only reveal different ligand recognition and activation mechanisms, but also provide a structural basis for drug optimization and design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30445.map.gz emd_30445.map.gz | 43.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30445-v30.xml emd-30445-v30.xml emd-30445.xml emd-30445.xml | 13.5 KB 13.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_30445.png emd_30445.png | 125.9 KB | ||
| Filedesc metadata |  emd-30445.cif.gz emd-30445.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30445 http://ftp.pdbj.org/pub/emdb/structures/EMD-30445 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30445 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30445 | HTTPS FTP |
-Related structure data
| Related structure data |  7cr2MC  7cr0C  7cr1C  7cr3C  7cr4C  7cr7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30445.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30445.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.014 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : voltage-gated potassium channel KCNQ2
| Entire | Name: voltage-gated potassium channel KCNQ2 |
|---|---|
| Components |
|
-Supramolecule #1: voltage-gated potassium channel KCNQ2
| Supramolecule | Name: voltage-gated potassium channel KCNQ2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Potassium voltage-gated channel subfamily KQT member 2
| Macromolecule | Name: Potassium voltage-gated channel subfamily KQT member 2 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 73.627812 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAGKPPKRNA FYRKLQNFLY NVLERPRGWA FIYHAYVFLL VFSCLVLSVF STIKEYEKSS EGALYILEIV TIVVFGVEYF VRIWAAGCC CRYRGWRGRL KFARKPFCVI DIMVLIASIA VLAAGSQGNV FATSALRSLR FLQILRMIRM DRRGGTWKLL G SVVYAHSK ...String: MAGKPPKRNA FYRKLQNFLY NVLERPRGWA FIYHAYVFLL VFSCLVLSVF STIKEYEKSS EGALYILEIV TIVVFGVEYF VRIWAAGCC CRYRGWRGRL KFARKPFCVI DIMVLIASIA VLAAGSQGNV FATSALRSLR FLQILRMIRM DRRGGTWKLL G SVVYAHSK ELVTAWYIGF LCLILASFLV YLAEKGENDH FDTYADALWW GLITLTTIGY GDKYPQTWNG RLLAATFTLI GV SFFALPA GILGSGFALK VQEQHRQKHF EKRRNPAAGL IQSAWRFYAT NLSRTDLHST WQYYERTVTV PMYSSQTQTY GAS RLIPPL NQLELLRNLK SKSGLAFRKD PPPEPSPSKG SPCRGPLCGC CPGRSSQKVS LKDRVFSSPR GVAAKGKGSP QAQT VRRSP SADQSLEDSP SKVPKSWSFG DRSRARQAFR IKGAASRQNS EEASLPGEDI VDDKSCPCEF VTEDLTPGLK VSIRA VCVM RFLVSKRKFK ESLRPYDVMD VIEQYSAGHL DMLSRIKSLQ SRVDQIVGRG PAITDKDRTK GPAEAELPED PSMMGR LGK VEKQVLSMEK KLDFLVNIYM QRMGIPPTET EAYFGAKEPE PAPPYHSPED SREHVDRHGC IVKIVRSSSS TGQKNFS VE GGSSGGWSHP QFEK UniProtKB: Potassium voltage-gated channel subfamily KQT member 2 |
-Macromolecule #2: ethyl N-[2-azanyl-4-[(4-fluorophenyl)methylamino]phenyl]carbamate
| Macromolecule | Name: ethyl N-[2-azanyl-4-[(4-fluorophenyl)methylamino]phenyl]carbamate type: ligand / ID: 2 / Number of copies: 4 / Formula: FBX |
|---|---|
| Molecular weight | Theoretical: 303.331 Da |
| Chemical component information | 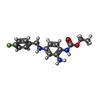 ChemComp-FBX: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.556 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















