+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-30310 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Human RAD51 post-synaptic complexes mutant (V273P, D274G) | ||||||||||||
 マップデータ マップデータ | |||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | meitotic homologous recombination / DNA repair / ATPase / RECOMBINATION | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報presynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / chromosome organization involved in meiotic cell cycle / DNA recombinase assembly / cellular response to camptothecin / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin ...presynaptic intermediate filament cytoskeleton / response to glucoside / mitotic recombination-dependent replication fork processing / chromosome organization involved in meiotic cell cycle / DNA recombinase assembly / cellular response to camptothecin / telomere maintenance via telomere lengthening / double-strand break repair involved in meiotic recombination / nuclear ubiquitin ligase complex / cellular response to cisplatin / DNA strand invasion / cellular response to hydroxyurea / mitotic recombination / DNA strand exchange activity / replication-born double-strand break repair via sister chromatid exchange / lateral element / regulation of DNA damage checkpoint / Impaired BRCA2 binding to PALB2 / telomere maintenance via recombination / reciprocal meiotic recombination / single-stranded DNA helicase activity / Homologous DNA Pairing and Strand Exchange / Defective homologous recombination repair (HRR) due to BRCA1 loss of function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA1 binding function / Defective HDR through Homologous Recombination Repair (HRR) due to PALB2 loss of BRCA2/RAD51/RAD51C binding function / Resolution of D-loop Structures through Synthesis-Dependent Strand Annealing (SDSA) / Resolution of D-loop Structures through Holliday Junction Intermediates / HDR through Single Strand Annealing (SSA) / regulation of double-strand break repair via homologous recombination / ATP-dependent DNA damage sensor activity / nuclear chromosome / Impaired BRCA2 binding to RAD51 / Transcriptional Regulation by E2F6 / replication fork processing / Presynaptic phase of homologous DNA pairing and strand exchange / response to X-ray / ATP-dependent activity, acting on DNA / interstrand cross-link repair / condensed chromosome / DNA polymerase binding / cellular response to ionizing radiation / condensed nuclear chromosome / meiotic cell cycle / double-strand break repair via homologous recombination / male germ cell nucleus / HDR through Homologous Recombination (HRR) / cellular response to gamma radiation / PML body / Meiotic recombination / response to toxic substance / site of double-strand break / single-stranded DNA binding / double-stranded DNA binding / DNA recombination / chromosome, telomeric region / mitochondrial matrix / response to xenobiotic stimulus / DNA repair / centrosome / DNA damage response / chromatin binding / chromatin / nucleolus / perinuclear region of cytoplasm / enzyme binding / protein-containing complex / ATP hydrolysis activity / mitochondrion / nucleoplasm / ATP binding / identical protein binding / nucleus / cytosol / cytoplasm 類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | ||||||||||||
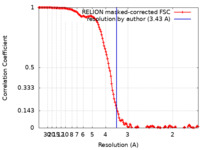
| 手法 | らせん対称体再構成法 / クライオ電子顕微鏡法 / 解像度: 3.43 Å | ||||||||||||
 データ登録者 データ登録者 | Chi HY / Ho MC / Tsai MD / Luo SC / Yeh HY | ||||||||||||
| 資金援助 |  台湾, 3件 台湾, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2021 ジャーナル: Nat Commun / 年: 2021タイトル: Identification of fidelity-governing factors in human recombinases DMC1 and RAD51 from cryo-EM structures. 著者: Shih-Chi Luo / Hsin-Yi Yeh / Wei-Hsuan Lan / Yi-Min Wu / Cheng-Han Yang / Hao-Yen Chang / Guan-Chin Su / Chia-Yi Lee / Wen-Jin Wu / Hung-Wen Li / Meng-Chiao Ho / Peter Chi / Ming-Daw Tsai /  要旨: Both high-fidelity and mismatch-tolerant recombination, catalyzed by RAD51 and DMC1 recombinases, respectively, are indispensable for genomic integrity. Here, we use cryo-EM, MD simulation and ...Both high-fidelity and mismatch-tolerant recombination, catalyzed by RAD51 and DMC1 recombinases, respectively, are indispensable for genomic integrity. Here, we use cryo-EM, MD simulation and functional analysis to elucidate the structural basis for the mismatch tolerance of DMC1. Structural analysis of DMC1 presynaptic and postsynaptic complexes suggested that the lineage-specific Loop 1 Gln244 (Met243 in RAD51) may help stabilize DNA backbone, whereas Loop 2 Pro274 and Gly275 (Val273/Asp274 in RAD51) may provide an open "triplet gate" for mismatch tolerance. In support, DMC1-Q244M displayed marked increase in DNA dynamics, leading to unobservable DNA map. MD simulation showed highly dispersive mismatched DNA ensemble in RAD51 but well-converged DNA in DMC1 and RAD51-V273P/D274G. Replacing Loop 1 or Loop 2 residues in DMC1 with RAD51 counterparts enhanced DMC1 fidelity, while reciprocal mutations in RAD51 attenuated its fidelity. Our results show that three Loop 1/Loop 2 residues jointly enact contrasting fidelities of DNA recombinases. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_30310.map.gz emd_30310.map.gz | 202.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-30310-v30.xml emd-30310-v30.xml emd-30310.xml emd-30310.xml | 16.8 KB 16.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_30310_fsc.xml emd_30310_fsc.xml | 13.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_30310.png emd_30310.png | 106 KB | ||
| Filedesc metadata |  emd-30310.cif.gz emd-30310.cif.gz | 6.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30310 http://ftp.pdbj.org/pub/emdb/structures/EMD-30310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30310 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_30310_validation.pdf.gz emd_30310_validation.pdf.gz | 603.7 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_30310_full_validation.pdf.gz emd_30310_full_validation.pdf.gz | 603.2 KB | 表示 | |
| XML形式データ |  emd_30310_validation.xml.gz emd_30310_validation.xml.gz | 13.4 KB | 表示 | |
| CIF形式データ |  emd_30310_validation.cif.gz emd_30310_validation.cif.gz | 18.1 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30310 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30310 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30310 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30310 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_30310.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_30310.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : RAD51-dsDNA filament (V273P D274G mutant)
| 全体 | 名称: RAD51-dsDNA filament (V273P D274G mutant) |
|---|---|
| 要素 |
|
-超分子 #1: RAD51-dsDNA filament (V273P D274G mutant)
| 超分子 | 名称: RAD51-dsDNA filament (V273P D274G mutant) / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1-#3 |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: DNA repair protein RAD51 homolog 1
| 分子 | 名称: DNA repair protein RAD51 homolog 1 / タイプ: protein_or_peptide / ID: 1 / コピー数: 3 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 36.949074 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MAMQMQLEAN ADTSVEEESF GPQPISRLEQ CGINANDVKK LEEAGFHTVE AVAYAPKKEL INIKGISEAK ADKILAEAAK LVPMGFTTA TEFHQRRSEI IQITTGSKEL DKLLQGGIET GSITEMFGEF RTGKTQICHT LAVTCQLPID RGGGEGKAMY I DTEGTFRP ...文字列: MAMQMQLEAN ADTSVEEESF GPQPISRLEQ CGINANDVKK LEEAGFHTVE AVAYAPKKEL INIKGISEAK ADKILAEAAK LVPMGFTTA TEFHQRRSEI IQITTGSKEL DKLLQGGIET GSITEMFGEF RTGKTQICHT LAVTCQLPID RGGGEGKAMY I DTEGTFRP ERLLAVAERY GLSGSDVLDN VAYARAFNTD HQTQLLYQAS AMMVESRYAL LIVDSATALY RTDYSGRGEL SA RQMHLAR FLRMLLRLAD EFGVAVVITN QVVAQPGGAA MFAADPKKPI GGNIIAHAST TRLYLRKGRG ETRICKIYDS PCL PEAEAM FAINADGVGD AKD UniProtKB: DNA repair protein RAD51 homolog 1 |
-分子 #2: DNA (5'-D(P*TP*TP*TP*TP*TP*TP*TP*TP*T)-3')
| 分子 | 名称: DNA (5'-D(P*TP*TP*TP*TP*TP*TP*TP*TP*T)-3') / タイプ: dna / ID: 2 詳細: Authors know the sequence of chains D/E. Chain D: TCA GCT GTT GCC CGT. Chain E: ACG GGC AAC AGC TGA. Since the DNA sequence could not be revealed by the helical reconstruction method, authors ...詳細: Authors know the sequence of chains D/E. Chain D: TCA GCT GTT GCC CGT. Chain E: ACG GGC AAC AGC TGA. Since the DNA sequence could not be revealed by the helical reconstruction method, authors have used poly-T for chain D, poly-A for chain E respectively for model building. コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 2.692778 KDa |
| 配列 | 文字列: (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) |
-分子 #3: DNA (5'-D(P*AP*AP*AP*AP*AP*AP*AP*AP*A)-3')
| 分子 | 名称: DNA (5'-D(P*AP*AP*AP*AP*AP*AP*AP*AP*A)-3') / タイプ: dna / ID: 3 詳細: Authors know the sequence of chains D/E. Chain D: TCA GCT GTT GCC CGT. Chain E: ACG GGC AAC AGC TGA. Since the DNA sequence could not be revealed by the helical reconstruction method, authors ...詳細: Authors know the sequence of chains D/E. Chain D: TCA GCT GTT GCC CGT. Chain E: ACG GGC AAC AGC TGA. Since the DNA sequence could not be revealed by the helical reconstruction method, authors have used poly-T for chain D, poly-A for chain E respectively for model building. コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 2.773904 KDa |
| 配列 | 文字列: (DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA)(DA) |
-分子 #4: CALCIUM ION
| 分子 | 名称: CALCIUM ION / タイプ: ligand / ID: 4 / コピー数: 3 / 式: CA |
|---|---|
| 分子量 | 理論値: 40.078 Da |
-分子 #5: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| 分子 | 名称: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / タイプ: ligand / ID: 5 / コピー数: 3 / 式: ANP |
|---|---|
| 分子量 | 理論値: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | らせん対称体再構成法 |
| 試料の集合状態 | filament |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 / 構成要素 - 濃度: 25.0 mM / 構成要素 - 式: Tris-HCl 詳細: 25 mM Tris-HCl, pH 7.5, 50 mM KCl and 1 mM dithiothreitol) containing 2 mM AMP-PNP and 5 mM CaCl2 |
|---|---|
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: COPPER / メッシュ: 200 / 支持フィルム - 材質: GRAPHENE OXIDE / 支持フィルム - トポロジー: CONTINUOUS / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 15 sec. |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 295 K / 装置: FEI VITROBOT MARK IV 詳細: The grids were blotted for 1 sec at 22 degree C with 100% relative humidity and plunge-frozen in liquid ethane cooled by liquid nitrogen using a Vitrobot Mark IV (Thermo Fisher).. |
| 詳細 | protein sample were applied onto a pre-glow-discharged graphene-oxide coated Quantifoil holey carbon grid (1.2/1.3, 200 mesh) using published protocol |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| アライメント法 | Coma free - Residual tilt: 10.0 mrad |
| 特殊光学系 | エネルギーフィルター - 名称: GIF Quantum LS / エネルギーフィルター - スリット幅: 30 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 検出モード: COUNTING / デジタル化 - サイズ - 横: 3838 pixel / デジタル化 - サイズ - 縦: 3710 pixel / デジタル化 - 画像ごとのフレーム数: 1-50 / 撮影したグリッド数: 1 / 平均露光時間: 5.0 sec. / 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 70.0 µm / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 倍率(公称値): 165000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)