[English] 日本語
 Yorodumi
Yorodumi- EMDB-2370: The Respiratory Syncytial Virus nucleoprotein-RNA complex forms a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2370 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The Respiratory Syncytial Virus nucleoprotein-RNA complex forms a left-handed helical nucleocapsid. | |||||||||
 Map data Map data | Subtomogram average of purified recombinant measles virus nucleocapsid | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Respiratory Syncytial Virus / nucleocapsid / cryo-electron tomography / sub-tomogram averaging | |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / negative staining | |||||||||
 Authors Authors | Bakker SE / Duquerroy S / Galloux M / Loney C / Conner E / Eleouet JF / Rey FA / Bhella D | |||||||||
 Citation Citation |  Journal: J Gen Virol / Year: 2013 Journal: J Gen Virol / Year: 2013Title: The respiratory syncytial virus nucleoprotein-RNA complex forms a left-handed helical nucleocapsid. Authors: Saskia E Bakker / Stéphane Duquerroy / Marie Galloux / Colin Loney / Edward Conner / Jean-François Eléouët / Félix A Rey / David Bhella /   Abstract: Respiratory syncytial virus (RSV) is an important human pathogen. Its nucleocapsid (NC), which comprises the negative sense RNA viral genome coated by the viral nucleoprotein N, is a critical ...Respiratory syncytial virus (RSV) is an important human pathogen. Its nucleocapsid (NC), which comprises the negative sense RNA viral genome coated by the viral nucleoprotein N, is a critical assembly that serves as template for both mRNA synthesis and genome replication. We have previously described the X-ray structure of an NC-like structure: a decameric ring formed of N-RNA that mimics one turn of the helical NC. In the absence of experimental data we had hypothesized that the NC helix would be right-handed, as the N-N contacts in the ring appeared to more easily adapt to that conformation. We now unambiguously show that the RSV NC is a left-handed helix. We further show that the contacts in the ring can be distorted to maintain key N-N-protein interactions in a left-handed helix, and discuss the implications of the resulting atomic model of the helical NC for viral replication and transcription. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2370.map.gz emd_2370.map.gz | 391.1 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2370-v30.xml emd-2370-v30.xml emd-2370.xml emd-2370.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2370.png EMD-2370.png | 55.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2370 http://ftp.pdbj.org/pub/emdb/structures/EMD-2370 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2370 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2370 | HTTPS FTP |
-Validation report
| Summary document |  emd_2370_validation.pdf.gz emd_2370_validation.pdf.gz | 206.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2370_full_validation.pdf.gz emd_2370_full_validation.pdf.gz | 205.3 KB | Display | |
| Data in XML |  emd_2370_validation.xml.gz emd_2370_validation.xml.gz | 4.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2370 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2370 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2370 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2370 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2370.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2370.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
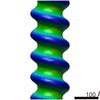
| Annotation | Subtomogram average of purified recombinant measles virus nucleocapsid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Purified recombinant measles virus nucleocapsids.
| Entire | Name: Purified recombinant measles virus nucleocapsids. |
|---|---|
| Components |
|
-Supramolecule #1000: Purified recombinant measles virus nucleocapsids.
| Supramolecule | Name: Purified recombinant measles virus nucleocapsids. / type: sample / ID: 1000 / Oligomeric state: variable helices / Number unique components: 1 |
|---|
-Supramolecule #1: Measles virus
| Supramolecule | Name: Measles virus / type: virus / ID: 1 / Name.synonym: MeV / Details: Purified recombinant nucleocapsid / NCBI-ID: 11234 / Sci species name: Measles virus / Virus type: OTHER / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No / Syn species name: MeV |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Host system | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 10 mM EDTA |
|---|---|
| Staining | Type: NEGATIVE Details: Protein preparations were mixed with 10 nm colloidal gold and loaded onto freshly glow-discharged perforated C-flat carbon films (CF-22-2C, Protochips Inc.). Grids were then washed in 0.1% ...Details: Protein preparations were mixed with 10 nm colloidal gold and loaded onto freshly glow-discharged perforated C-flat carbon films (CF-22-2C, Protochips Inc.). Grids were then washed in 0.1% trehalose, and stained using 5% ammonium molybdate (pH 7.4) and 0.1% trehalose, drained and allowed to dry in air |
| Grid | Details: Glow-discharged perforated C-flat carbon films (CF-22-2C, Protochips Inc.) |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2200FSC |
|---|---|
| Temperature | Min: 97 K / Max: 293 K |
| Alignment procedure | Legacy - Astigmatism: Corrected at 100k times mag in digital micrograph |
| Specialist optics | Energy filter - Name: Jeol / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 30.0 eV |
| Details | Room temperature stained grids imaged at LN2 temperatures. |
| Date | Jul 15, 2010 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Number real images: 7 / Average electron dose: 131 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 40000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN / Tilt series - Axis1 - Min angle: -60 ° / Tilt series - Axis1 - Max angle: 60 ° |
- Image processing
Image processing
| Details | Subtomograms were manually picked. A starting model was obtained by manually aligning 100 subtomograms, all subtomograms were aligned to this starting model. To mitigate missing wedge artefacts, the orientations around the helical axis were randomised before further alignment. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Software - Name: Etomo, Dynamo / Details: Subtomograms were picked from nine tomograms. / Number subtomograms used: 911 |
| Final 3D classification | Number classes: 1 |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)