+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-23665 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structural basis for SARS-CoV-2 envelope protein in recognition of human cell junction protein PALS1 | |||||||||
 マップデータ マップデータ | Cryosparc non-uniform refinement without sharpening | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | SARS-CoV-2 envelope protein / PDZ-binding motif / complex / pathogen-host interaction / CELL ADHESION-VIRAL PROTEIN complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報disruption of cellular anatomical structure in another organism / protein localization to myelin sheath abaxonal region / SARS-CoV-1 targets PDZ proteins in cell-cell junction / viral budding from Golgi membrane / establishment or maintenance of polarity of embryonic epithelium / myelin assembly / morphogenesis of an epithelial sheet / Tight junction interactions / SARS-CoV-2 targets PDZ proteins in cell-cell junction / cytoplasmic capsid assembly ...disruption of cellular anatomical structure in another organism / protein localization to myelin sheath abaxonal region / SARS-CoV-1 targets PDZ proteins in cell-cell junction / viral budding from Golgi membrane / establishment or maintenance of polarity of embryonic epithelium / myelin assembly / morphogenesis of an epithelial sheet / Tight junction interactions / SARS-CoV-2 targets PDZ proteins in cell-cell junction / cytoplasmic capsid assembly / regulation of transforming growth factor beta receptor signaling pathway / lateral loop / myelin sheath adaxonal region / Regulation of gap junction activity / Schmidt-Lanterman incisure / establishment or maintenance of epithelial cell apical/basal polarity / peripheral nervous system myelin maintenance / apical junction complex / generation of neurons / central nervous system neuron development / endoplasmic reticulum-Golgi intermediate compartment / host cell Golgi membrane / bicellular tight junction / Maturation of protein E / endoplasmic reticulum-Golgi intermediate compartment membrane / monoatomic ion channel activity / protein localization to plasma membrane / adherens junction / cerebral cortex development / gene expression / perikaryon / Translation of Structural Proteins / Virion Assembly and Release / Induction of Cell-Cell Fusion / structural constituent of virion / Attachment and Entry / apical plasma membrane / protein domain specific binding / axon / apoptotic process / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / Golgi apparatus / protein-containing complex / extracellular exosome / ATP binding / identical protein binding / membrane / plasma membrane / cytoplasm 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  | |||||||||
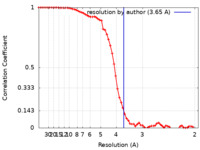
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.65 Å | |||||||||
 データ登録者 データ登録者 | Liu Q / Chai J | |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2021 ジャーナル: Nat Commun / 年: 2021タイトル: Structural basis for SARS-CoV-2 envelope protein recognition of human cell junction protein PALS1. 著者: Jin Chai / Yuanheng Cai / Changxu Pang / Liguo Wang / Sean McSweeney / John Shanklin / Qun Liu /  要旨: The COVID-19 pandemic, caused by the SARS-CoV-2 virus, has created global health and economic emergencies. SARS-CoV-2 viruses promote their own spread and virulence by hijacking human proteins, which ...The COVID-19 pandemic, caused by the SARS-CoV-2 virus, has created global health and economic emergencies. SARS-CoV-2 viruses promote their own spread and virulence by hijacking human proteins, which occurs through viral protein recognition of human targets. To understand the structural basis for SARS-CoV-2 viral-host protein recognition, here we use cryo-electron microscopy (cryo-EM) to determine a complex structure of the human cell junction protein PALS1 and SARS-CoV-2 viral envelope (E) protein. Our reported structure shows that the E protein C-terminal DLLV motif recognizes a pocket formed exclusively by hydrophobic residues from the PDZ and SH3 domains of PALS1. Our structural analysis provides an explanation for the observation that the viral E protein recruits PALS1 from lung epithelial cell junctions. In addition, our structure provides novel targets for peptide- and small-molecule inhibitors that could block the PALS1-E interactions to reduce E-mediated virulence. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_23665.map.gz emd_23665.map.gz | 31.8 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-23665-v30.xml emd-23665-v30.xml emd-23665.xml emd-23665.xml | 9.9 KB 9.9 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_23665_fsc.xml emd_23665_fsc.xml | 6.4 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_23665.png emd_23665.png | 152.4 KB | ||
| Filedesc metadata |  emd-23665.cif.gz emd-23665.cif.gz | 5.2 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23665 http://ftp.pdbj.org/pub/emdb/structures/EMD-23665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23665 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_23665_validation.pdf.gz emd_23665_validation.pdf.gz | 474.8 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_23665_full_validation.pdf.gz emd_23665_full_validation.pdf.gz | 474.3 KB | 表示 | |
| XML形式データ |  emd_23665_validation.xml.gz emd_23665_validation.xml.gz | 9.8 KB | 表示 | |
| CIF形式データ |  emd_23665_validation.cif.gz emd_23665_validation.cif.gz | 12.5 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23665 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23665 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23665 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23665 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_23665.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_23665.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Cryosparc non-uniform refinement without sharpening | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.684 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
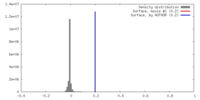
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Complex structure of SARS-CoV-2 envelope protein Ec18 and human P...
| 全体 | 名称: Complex structure of SARS-CoV-2 envelope protein Ec18 and human PALS1 PSG domains |
|---|---|
| 要素 |
|
-超分子 #1: Complex structure of SARS-CoV-2 envelope protein Ec18 and human P...
| 超分子 | 名称: Complex structure of SARS-CoV-2 envelope protein Ec18 and human PALS1 PSG domains タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: MAGUK p55 subfamily member 5
| 分子 | 名称: MAGUK p55 subfamily member 5 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 44.780504 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: SNAPITDERV YESIGQYGGE TVKIVRIEKA RDIPLGATVR NEMDSVIISR IVKGGAAEKS GLLHEGDEVL EINGIEIRGK DVNEVFDLL SDMHGTLTFV LIPSQQIKPP PAKETVIHVK AHFDYDPSDD PYVPCRELGL SFQKGDILHV ISQEDPNWWQ A YREGDEDN ...文字列: SNAPITDERV YESIGQYGGE TVKIVRIEKA RDIPLGATVR NEMDSVIISR IVKGGAAEKS GLLHEGDEVL EINGIEIRGK DVNEVFDLL SDMHGTLTFV LIPSQQIKPP PAKETVIHVK AHFDYDPSDD PYVPCRELGL SFQKGDILHV ISQEDPNWWQ A YREGDEDN QPLAGLVPGK EEILTYEEMS LYHQPANRKR PIILIGPQNC GQNELRQRLM NKEKDRFASA VPHTTRSRRD QE VAGRDYH FVSRQAFEAD IAAGKFIEHG EFEKNLYGTS IDSVRQVINS GKICLLSLRT QSLKTLRNSD LKPYIIFIAP PSQ ERLRAL LAKEGKNPKP EELREIIEKT REMEQNNGHY FDTAIVNSDL DKAYQELLRL INKLDTEPQW VPSTWLR UniProtKB: Protein PALS1, Protein PALS1 |
-分子 #2: Envelope small membrane protein
| 分子 | 名称: Envelope small membrane protein / タイプ: protein_or_peptide / ID: 2 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 2.062395 KDa |
| 配列 | 文字列: VYSRVKNLNS SRVPDLLV UniProtKB: Envelope small membrane protein |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) 平均電子線量: 64.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)