+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21959 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Colicin E1 fragment in nanodisc-embedded TolC | ||||||||||||
 Map data Map data | asymmetric map of colicin E1 bound to TolC in nanodisc; cropped, sharpened, and resampled | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | antibiotic efflux / bacteriocin / TRANSPORT PROTEIN / ANTIMICROBIAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of ion transmembrane transporter activity / MacAB-TolC complex / enterobactin transport / pore-forming activity / bile acid transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / efflux pump complex / periplasmic side of plasma membrane / xenobiotic detoxification by transmembrane export across the plasma membrane / bile acid and bile salt transport ...negative regulation of ion transmembrane transporter activity / MacAB-TolC complex / enterobactin transport / pore-forming activity / bile acid transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / efflux pump complex / periplasmic side of plasma membrane / xenobiotic detoxification by transmembrane export across the plasma membrane / bile acid and bile salt transport / porin activity / efflux transmembrane transporter activity / monoatomic ion transmembrane transport / cell outer membrane / response to organic cyclic compound / response to toxic substance / monoatomic ion channel activity / outer membrane-bounded periplasmic space / defense response to Gram-negative bacterium / killing of cells of another organism / transmembrane transporter binding / response to xenobiotic stimulus / response to antibiotic / identical protein binding / membrane / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |    | ||||||||||||
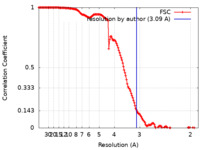
| Method | single particle reconstruction / cryo EM / Resolution: 3.09 Å | ||||||||||||
 Authors Authors | Kaelber JT / Budiardjo SJ | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Colicin E1 opens its hinge to plug TolC. Authors: S Jimmy Budiardjo / Jacqueline J Stevens / Anna L Calkins / Ayotunde P Ikujuni / Virangika K Wimalasena / Emre Firlar / David A Case / Julie S Biteen / Jason T Kaelber / Joanna S G Slusky /  Abstract: The double membrane architecture of Gram-negative bacteria forms a barrier that is impermeable to most extracellular threats. Bacteriocin proteins evolved to exploit the accessible, surface-exposed ...The double membrane architecture of Gram-negative bacteria forms a barrier that is impermeable to most extracellular threats. Bacteriocin proteins evolved to exploit the accessible, surface-exposed proteins embedded in the outer membrane to deliver cytotoxic cargo. Colicin E1 is a bacteriocin produced by, and lethal to, that hijacks the outer membrane proteins (OMPs) TolC and BtuB to enter the cell. Here, we capture the colicin E1 translocation domain inside its membrane receptor, TolC, by high-resolution cryo-electron microscopy to obtain the first reported structure of a bacteriocin bound to TolC. Colicin E1 binds stably to TolC as an open hinge through the TolC pore-an architectural rearrangement from colicin E1's unbound conformation. This binding is stable in live cells as indicated by single-molecule fluorescence microscopy. Finally, colicin E1 fragments binding to TolC plug the channel, inhibiting its native efflux function as an antibiotic efflux pump, and heightening susceptibility to three antibiotic classes. In addition to demonstrating that these protein fragments are useful starting points for developing novel antibiotic potentiators, this method could be expanded to other colicins to inhibit other OMP functions. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21959.map.gz emd_21959.map.gz | 52.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21959-v30.xml emd-21959-v30.xml emd-21959.xml emd-21959.xml | 24.4 KB 24.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21959_fsc.xml emd_21959_fsc.xml | 15.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_21959.png emd_21959.png | 64.1 KB | ||
| Masks |  emd_21959_msk_1.map emd_21959_msk_1.map | 282.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-21959.cif.gz emd-21959.cif.gz | 7.1 KB | ||
| Others |  emd_21959_additional_1.map.gz emd_21959_additional_1.map.gz emd_21959_half_map_1.map.gz emd_21959_half_map_1.map.gz emd_21959_half_map_2.map.gz emd_21959_half_map_2.map.gz | 145.9 MB 262 MB 262 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21959 http://ftp.pdbj.org/pub/emdb/structures/EMD-21959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21959 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21959 | HTTPS FTP |
-Validation report
| Summary document |  emd_21959_validation.pdf.gz emd_21959_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21959_full_validation.pdf.gz emd_21959_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_21959_validation.xml.gz emd_21959_validation.xml.gz | 22.5 KB | Display | |
| Data in CIF |  emd_21959_validation.cif.gz emd_21959_validation.cif.gz | 28.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21959 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21959 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21959 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21959 | HTTPS FTP |
-Related structure data
| Related structure data |  6wxhMC  6wxiC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21959.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21959.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | asymmetric map of colicin E1 bound to TolC in nanodisc; cropped, sharpened, and resampled | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_21959_msk_1.map emd_21959_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
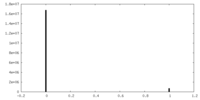
| Density Histograms |
-Additional map: C3-symmetrized refinement of colicin E1 bound to TolC...
| File | emd_21959_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C3-symmetrized refinement of colicin E1 bound to TolC in nanodisc; sharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
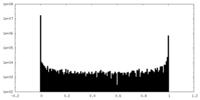
| Density Histograms |
-Half map: unmasked, unfiltered cryoSPARC half-map B of colicin E1...
| File | emd_21959_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unmasked, unfiltered cryoSPARC half-map B of colicin E1 bound to TolC in nanodisc | ||||||||||||
| Projections & Slices |
| ||||||||||||
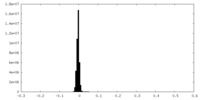
| Density Histograms |
-Half map: unmasked, unfiltered cryoSPARC half-map A of colicin E1...
| File | emd_21959_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unmasked, unfiltered cryoSPARC half-map A of colicin E1 bound to TolC in nanodisc | ||||||||||||
| Projections & Slices |
| ||||||||||||
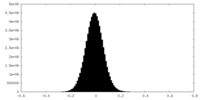
| Density Histograms |
- Sample components
Sample components
-Entire : Colicin E1 fragment colE1-T inside nanodisc-embedded TolC
| Entire | Name: Colicin E1 fragment colE1-T inside nanodisc-embedded TolC |
|---|---|
| Components |
|
-Supramolecule #1: Colicin E1 fragment colE1-T inside nanodisc-embedded TolC
| Supramolecule | Name: Colicin E1 fragment colE1-T inside nanodisc-embedded TolC type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 21.44974 KDa |
-Supramolecule #2: Outer membrane protein TolC
| Supramolecule | Name: Outer membrane protein TolC / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Colicin E1 fragment colE1-T
| Supramolecule | Name: Colicin E1 fragment colE1-T / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Outer membrane protein TolC
| Macromolecule | Name: Outer membrane protein TolC / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.783355 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKKLLPILIG LSLSGFSSLS QAENLMQVYQ QARLSNPELR KSAADRDAAF EKINEARSPL LPQLGLGADY TYSNGYRDAN GINSNATSA SLQLTQSIFD MSKWRALTLQ EKAAGIQDVT YQTDQQTLIL NTATAYFNVL NAIDVLSYTQ AQKEAIYRQL D QTTQRFNV ...String: MKKLLPILIG LSLSGFSSLS QAENLMQVYQ QARLSNPELR KSAADRDAAF EKINEARSPL LPQLGLGADY TYSNGYRDAN GINSNATSA SLQLTQSIFD MSKWRALTLQ EKAAGIQDVT YQTDQQTLIL NTATAYFNVL NAIDVLSYTQ AQKEAIYRQL D QTTQRFNV GLVAITDVQN ARAQYDTVLA NEVTARNNLD NAVEQLRQIT GNYYPELAAL NVENFKTDKP QPVNALLKEA EK RNLSLLQ ARLSQDLARE QIRQAQDGHL PTLDLTASTG ISDTSYSGSK TRGAAGTQYD DSNMGQNKVG LSFSLPIYQG GMV NSQVKQ AQYNFVGASE QLESAHRSVV QTVRSSFNNI NASISSINAY KQAVVSAQSS LDAMEAGYSV GTRTIVDVLD ATTT LYNAK QELANARYNY LINQLNIKSA LGTLNEQDLL ALNNALSKPV STNPENVAPQ TPEQNAIADG YAPDSPAPVV QQTSA RTTT SNGHNPFRN UniProtKB: Outer membrane protein TolC |
-Macromolecule #2: Colicin-E1
| Macromolecule | Name: Colicin-E1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.491807 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: METAVAYYKD GVPYDDKGQV IITLLNGTPD GSGSGGGGGK GGSKSESSAA IHATAKWSTA QLKKTQAEQA ARAKAAAEAQ AKAKANRDA LTQRLKDIVN EALRHNASRT PSATELAHAN NAAMQAEDER LRLAKAEEKA RKEAEAAEKA FQEAEQRRKE I EREKAETE ...String: METAVAYYKD GVPYDDKGQV IITLLNGTPD GSGSGGGGGK GGSKSESSAA IHATAKWSTA QLKKTQAEQA ARAKAAAEAQ AKAKANRDA LTQRLKDIVN EALRHNASRT PSATELAHAN NAAMQAEDER LRLAKAEEKA RKEAEAAEKA FQEAEQRRKE I EREKAETE RQLKLAEAEE KRLAALSEEA KALEHHHHHH UniProtKB: Colicin-E1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: UltrAuFoil / Material: GOLD / Mesh: 200 / Support film - Material: GOLD / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Number real images: 10644 / Average exposure time: 6.0 sec. / Average electron dose: 35.96 e/Å2 / Details: 5 frames per second |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 20.0 µm / Calibrated defocus max: 2.1 µm / Calibrated defocus min: 0.4 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 205000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z
Z Y
Y X
X