+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2182 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron microscopy map of SOS1 antiporter | |||||||||
 Map data Map data | SOS1 antiporter class 2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | membrane protein / plant salt tolerance / protein structure | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.9 Å | |||||||||
 Authors Authors | Nunez Ramirez R / Sanchez Barrena MJ / Villalta I / Vega JF / Pardo JM / Quintero FJ / Martinez Salazar J / Albert A | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2012 Journal: J Mol Biol / Year: 2012Title: Structural insights on the plant salt-overly-sensitive 1 (SOS1) Na(+)/H(+) antiporter. Authors: Rafael Núñez-Ramírez / María José Sánchez-Barrena / Irene Villalta / Juan F Vega / Jose M Pardo / Francisco J Quintero / Javier Martinez-Salazar / Armando Albert /  Abstract: The Arabidopsisthaliana Na(+)/H(+) antiporter salt-overly-sensitive 1 (SOS1) is essential to maintain low intracellular levels of toxic Na(+) under salt stress. Available data show that the plant ...The Arabidopsisthaliana Na(+)/H(+) antiporter salt-overly-sensitive 1 (SOS1) is essential to maintain low intracellular levels of toxic Na(+) under salt stress. Available data show that the plant SOS2 protein kinase and its interacting activator, the SOS3 calcium-binding protein, function together in decoding calcium signals elicited by salt stress and regulating the phosphorylation state and the activity of SOS1. Molecular genetic studies have shown that the activation implies a domain reorganization of the antiporter cytosolic moiety, indicating that there is a clear relationship between function and molecular structure of the antiporter. To provide information on this issue, we have carried out in vivo and in vitro studies on the oligomerization state of SOS1. In addition, we have performed electron microscopy and single-particle reconstruction of negatively stained full-length and active SOS1. Our studies show that the protein is a homodimer that contains a membrane domain similar to that found in other antiporters of the family and an elongated, large, and structured cytosolic domain. Both the transmembrane (TM) and cytosolic moieties contribute to the dimerization of the antiporter. The close contacts between the TM and the cytosolic domains provide a link between regulation and transport activity of the antiporter. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2182.map.gz emd_2182.map.gz | 1.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2182-v30.xml emd-2182-v30.xml emd-2182.xml emd-2182.xml | 9.9 KB 9.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2182.tif emd_2182.tif | 2.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2182 http://ftp.pdbj.org/pub/emdb/structures/EMD-2182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2182 | HTTPS FTP |
-Validation report
| Summary document |  emd_2182_validation.pdf.gz emd_2182_validation.pdf.gz | 200.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2182_full_validation.pdf.gz emd_2182_full_validation.pdf.gz | 199.4 KB | Display | |
| Data in XML |  emd_2182_validation.xml.gz emd_2182_validation.xml.gz | 4.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2182 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2182 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2182 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2182 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2182.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2182.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SOS1 antiporter class 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
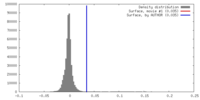
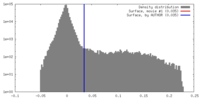
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Antiporter SOS1
| Entire | Name: Antiporter SOS1 |
|---|---|
| Components |
|
-Supramolecule #1000: Antiporter SOS1
| Supramolecule | Name: Antiporter SOS1 / type: sample / ID: 1000 Details: This sample is a membrane protein and is purified with lipids and detergents surrounding the membrane domain Oligomeric state: Dimer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 360 KDa / Theoretical: 254 KDa Method: Gel filtration, Electron microscopy image analysis, Dynamic light scattering |
-Macromolecule #1: Na+/H+ antiporter SOS1
| Macromolecule | Name: Na+/H+ antiporter SOS1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Oligomeric state: Dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 127 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 Details: 50 mM phosphate buffer pH 8.5, 300 mM NaCl, 10% glycerol, 0.05% DDM, 250 mM imidazole |
|---|---|
| Staining | Type: NEGATIVE Details: Purified SOS1 was adsorbed to glow-discharged carbon coated grids and stained with 2% uranyl formate |
| Grid | Details: 400 mesh carbon coated copper grid, glow discharged in moderate vacuum |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2100 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 120.000-150.000 times magnification and confirmed by inspection of the Fourier transform |
| Date | Nov 1, 2010 |
| Image recording | Category: CCD / Film or detector model: GATAN ORIUS SC200 (2k x 2k) / Number real images: 90 / Average electron dose: 15 e/Å2 / Bits/pixel: 14 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 20000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.0 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 20000 |
| Sample stage | Specimen holder model: JEOL |
- Image processing
Image processing
| Details | Single images of SOS1 were manually extracted using EMAN. 2D reference-free classification, 2D averaging, 3D classification and reconstruction were performed using maximum-likelihood methods implemented in XMIPP package. |
|---|---|
| CTF correction | Details: Each micrograph. Estimation using CTFFIND3 and correction using Bsoft |
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.9 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: XMIPP / Number images used: 6300 |
| Final two d classification | Number classes: 96 |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)