[English] 日本語
 Yorodumi
Yorodumi- EMDB-21678: GluN1b-GluN2B NMDA receptor in complex with GluN2B antagonist SDZ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21678 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | GluN1b-GluN2B NMDA receptor in complex with GluN2B antagonist SDZ 220-040, class 1 | |||||||||
 Map data Map data | B factor sharpened (-90) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NMDARs / Ligand-gated ion channels / METAL TRANSPORT / Ionotropic glutamate receptor / MEMBRANE PROTEIN / GluN2B antagonist | |||||||||
| Function / homology |  Function and homology information Function and homology informationcellular response to corticosterone stimulus / cellular response to magnesium starvation / sensory organ development / cellular response to curcumin / regulation of cAMP/PKA signal transduction / pons maturation / EPHB-mediated forward signaling / Assembly and cell surface presentation of NMDA receptors / regulation of cell communication / auditory behavior ...cellular response to corticosterone stimulus / cellular response to magnesium starvation / sensory organ development / cellular response to curcumin / regulation of cAMP/PKA signal transduction / pons maturation / EPHB-mediated forward signaling / Assembly and cell surface presentation of NMDA receptors / regulation of cell communication / auditory behavior / positive regulation of Schwann cell migration / sensitization / olfactory learning / response to other organism / response to hydrogen sulfide / dendritic branch / fear response / conditioned taste aversion / response to methylmercury / protein localization to postsynaptic membrane / regulation of ARF protein signal transduction / apical dendrite / regulation of respiratory gaseous exchange / response to manganese ion / transmitter-gated monoatomic ion channel activity / suckling behavior / interleukin-1 receptor binding / positive regulation of inhibitory postsynaptic potential / response to carbohydrate / cellular response to lipid / propylene metabolic process / response to glycine / cellular response to dsRNA / RAF/MAP kinase cascade / negative regulation of dendritic spine maintenance / response to amine / response to growth hormone / heterocyclic compound binding / neurotransmitter receptor complex / response to glycoside / Synaptic adhesion-like molecules / positive regulation of glutamate secretion / regulation of monoatomic cation transmembrane transport / NMDA glutamate receptor activity / voltage-gated monoatomic cation channel activity / NMDA selective glutamate receptor complex / glutamate binding / ligand-gated sodium channel activity / neuromuscular process / regulation of axonogenesis / calcium ion transmembrane import into cytosol / regulation of dendrite morphogenesis / male mating behavior / regulation of synapse assembly / protein heterotetramerization / response to morphine / small molecule binding / glycine binding / receptor clustering / startle response / positive regulation of reactive oxygen species biosynthetic process / parallel fiber to Purkinje cell synapse / regulation of MAPK cascade / monoatomic ion channel complex / behavioral response to pain / regulation of postsynaptic membrane potential / monoatomic cation transmembrane transport / action potential / response to electrical stimulus / positive regulation of calcium ion transport into cytosol / extracellularly glutamate-gated ion channel activity / cellular response to glycine / associative learning / positive regulation of dendritic spine maintenance / response to magnesium ion / Unblocking of NMDA receptors, glutamate binding and activation / regulation of neuronal synaptic plasticity / monoatomic cation transport / prepulse inhibition / glutamate receptor binding / detection of mechanical stimulus involved in sensory perception of pain / social behavior / ligand-gated monoatomic ion channel activity / response to mechanical stimulus / multicellular organismal response to stress / neuron development / phosphatase binding / long-term memory / postsynaptic density, intracellular component / behavioral fear response / monoatomic cation channel activity / synaptic cleft / response to fungicide / calcium ion homeostasis / positive regulation of synaptic transmission, glutamatergic / cellular response to manganese ion / glutamate-gated receptor activity / regulation of long-term synaptic depression / D2 dopamine receptor binding / glutamate-gated calcium ion channel activity Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.09 Å | |||||||||
 Authors Authors | Chou T / Tajima N | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Structural Basis of Functional Transitions in Mammalian NMDA Receptors. Authors: Tsung-Han Chou / Nami Tajima / Annabel Romero-Hernandez / Hiro Furukawa /  Abstract: Excitatory neurotransmission meditated by glutamate receptors including N-methyl-D-aspartate receptors (NMDARs) is pivotal to brain development and function. NMDARs are heterotetramers composed of ...Excitatory neurotransmission meditated by glutamate receptors including N-methyl-D-aspartate receptors (NMDARs) is pivotal to brain development and function. NMDARs are heterotetramers composed of GluN1 and GluN2 subunits, which bind glycine and glutamate, respectively, to activate their ion channels. Despite importance in brain physiology, the precise mechanisms by which activation and inhibition occur via subunit-specific binding of agonists and antagonists remain largely unknown. Here, we show the detailed patterns of conformational changes and inter-subunit and -domain reorientation leading to agonist-gating and subunit-dependent competitive inhibition by providing multiple structures in distinct ligand states at 4 Å or better. The structures reveal that activation and competitive inhibition by both GluN1 and GluN2 antagonists occur by controlling the tension of the linker between the ligand-binding domain and the transmembrane ion channel of the GluN2 subunit. Our results provide detailed mechanistic insights into NMDAR pharmacology, activation, and inhibition, which are fundamental to the brain physiology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21678.map.gz emd_21678.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21678-v30.xml emd-21678-v30.xml emd-21678.xml emd-21678.xml | 15.1 KB 15.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21678.png emd_21678.png | 67.1 KB | ||
| Filedesc metadata |  emd-21678.cif.gz emd-21678.cif.gz | 6.5 KB | ||
| Others |  emd_21678_additional.map.gz emd_21678_additional.map.gz | 59.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21678 http://ftp.pdbj.org/pub/emdb/structures/EMD-21678 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21678 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21678 | HTTPS FTP |
-Related structure data
| Related structure data |  6whwMC  6usuC  6usvC  6whrC  6whsC  6whtC  6whuC  6whvC  6whxC  6whyC  6wi0C  6wi1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21678.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21678.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | B factor sharpened (-90) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharpened map
| File | emd_21678_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : NMDA receptor GluN1b/2B functional ion channel complex
| Entire | Name: NMDA receptor GluN1b/2B functional ion channel complex |
|---|---|
| Components |
|
-Supramolecule #1: NMDA receptor GluN1b/2B functional ion channel complex
| Supramolecule | Name: NMDA receptor GluN1b/2B functional ion channel complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ionotropic glutamate receptor , NMDA receptor GluN1b
| Macromolecule | Name: Ionotropic glutamate receptor , NMDA receptor GluN1b / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 108.085633 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSTMHLLTFA LLFSCSFARA ASDPKIVNIG AVLSTRKHEQ MFREAVNQAN KRHGSWKIQL QATSVTHKPN AIQMALSVCE DLISSQVYA ILVSHPPTPN DHFTPTPVSY TAGFYRIPVL GLTTRMSIYS DKSIHLSFLR TVPPYSHQSS VWFEMMRVYN W NHIILLVS ...String: MSTMHLLTFA LLFSCSFARA ASDPKIVNIG AVLSTRKHEQ MFREAVNQAN KRHGSWKIQL QATSVTHKPN AIQMALSVCE DLISSQVYA ILVSHPPTPN DHFTPTPVSY TAGFYRIPVL GLTTRMSIYS DKSIHLSFLR TVPPYSHQSS VWFEMMRVYN W NHIILLVS DDHEGRAAQK RLETLLEERE SKSKKRNYEN LDQLSYDNKR GPKAEKVLQF DPGTKNVTAL LMEARELEAR VI ILSASED DAATVYRAAA MLDMTGSGYV WLVGEREISG NALRYAPDGI IGLQLINGKN ESAHISDAVG VVAQAVHELL EKE NITDPP RGCVGNTNIW KTGPLFKRVL MSSKYADGVT GRVEFNEDGD RKFAQYSIMN LQNRKLVQVG IYNGTHVIPN DRKI IWPGG ETEKPRGYQM STRLKIVTIH QEPFVYVKPT MSDGTCKEEF TVNGDPVKKV ICTGPNDTSP GSPRHTVPQC CYGFC IDLL IKLARTMQFT YEVHLVADGK FGTQERVQNS NKKEWNGMMG ELLSGQADMI VAPLTINNER AQYIEFSKPF KYQGLT ILV KKEIPRSTLD SFMQPFQSTL WLLVGLSVHV VAVMLYLLDR FSPFGRFKVN SQSESTDALT LSSAMWFSWG VLLNSGI GE GAPRSFSARI LGMVWAGFAM IIVASYTANL AAFLVLDRPE ERITGINDPR LRNPSDKFIY ATVKQSSVDI YFRRQVEL S TMYRHMEKHN YESAAEAIQA VRDNKLHAFI WDSAVLEFEA SQKCDLVTTG ELFFRSGFGI GMRKDSPWKQ QVSLSILKS HENGFMEDLD KTWVRYQECD SRSNAPATLT CENMAGVFML VAGGIVAGIF LIFIEIAYKR HKDARRKQMQ LAFAAVNVWR KNLQDRKSG RAEPDPKKKA TFRAITSTLA SSFKRRRSSK DTSTGGGRGA LQNQKDTVLP RRAIEREEGQ LQLCSRHRES |
-Macromolecule #2: Ionotropic glutamate receptor , NMDA receptor GluN2B
| Macromolecule | Name: Ionotropic glutamate receptor , NMDA receptor GluN2B / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 98.845859 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGTMRLFLLA VLFLFSFARA TGWSHPQFEK GGGSGGGSGG SAWSHPQFEK GALVPRGRSQ KSPPSIGIAV ILVGTSDEVA IKDAHEKDD FHHLSVVPRV ELVAMNETDP KSIITRICDL MSDRKIQGVV FADDTDQEAI AQILDFISAQ TLTPILGIHG G SSMIMADK ...String: MGTMRLFLLA VLFLFSFARA TGWSHPQFEK GGGSGGGSGG SAWSHPQFEK GALVPRGRSQ KSPPSIGIAV ILVGTSDEVA IKDAHEKDD FHHLSVVPRV ELVAMNETDP KSIITRICDL MSDRKIQGVV FADDTDQEAI AQILDFISAQ TLTPILGIHG G SSMIMADK DESSMFFQFG PSIEQQASVM LNIMEEYDWY IFSIVTTYFP GYQDFVNKIR STIENSFVGW ELEEVLLLDM SL DDGDSKI QNQLKKLQSP IILLYCTKEE ATYIFEVANS VGLTGYGYTW IVPSLVAGDT DTVPSEFPTG LISVSYDEWD YGL PARVRD GIAIITTAAS DMLSEHSFIP EPKSSCYNTH EKRIYQSNML NRYLINVTFE GRDLSFSEDG YQMHPKLVII LLNK ERKWE RVGKWKDKSL QMKYYVWPRM CPETEEQEDD HLSIVTLEEA PFVIVESVDP LSGTCMRNTV PCQKRIISEN KTDEE PGYI KKCCKGFCID ILKKISKSVK FTYDLYLVTN GKHGKKINGT WNGMIGEVVM KRAYMAVGSL TINEERSEVV DFSVPF IET GISVMVSRSN GTVSPSAFLE PFSACVWVMM FVMLLIVSAV AVFVFEYFSP VGYNRSLADG REPGGPSFTI GKAIWLL WG LVFNNSVPVQ NPKGTTSKIM VSVWAFFAVI FLASYTANLA AFMIQEEYVD QVSGLSDKKF QRPNDFSPPF RFGTVPNG S TERNIRNNYA EMHAYMGKFN QRGVDDALLS LKTGKLDAFI YDAAVLNYMA GRDEGCKLVT IGSGKVFAST GYGIAIQKD SGWKRQVDLA ILQLFGDGEM EELEALWLTG ICHNEKNEVM SSQLDIDNMA GVFYMLGAAM ALSLITFISE HLFYWQFRHS FMG |
-Macromolecule #3: (2S)-2-amino-3-[2',4'-dichloro-4-hydroxy-5-(phosphonomethyl)biphe...
| Macromolecule | Name: (2S)-2-amino-3-[2',4'-dichloro-4-hydroxy-5-(phosphonomethyl)biphenyl-3-yl]propanoic acid type: ligand / ID: 3 / Number of copies: 2 / Formula: QGP |
|---|---|
| Molecular weight | Theoretical: 420.181 Da |
| Chemical component information | 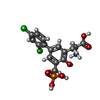 ChemComp-QGP: |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 295 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 64.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: Ab initio reconstruction |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 4.09 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cisTEM (ver. 1.0.0) / Number images used: 404527 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)