[English] 日本語
 Yorodumi
Yorodumi- EMDB-21385: Map of an acid-sensing ion channel in a resting state at high pH. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21385 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Map of an acid-sensing ion channel in a resting state at high pH. | |||||||||||||||
 Map data Map data | Map of an acid-sensing ion channel in a resting state at high pH. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||||||||
 Authors Authors | Yoder N / Jalali-Yazdi F / Gouaux E | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2020 Journal: J Struct Biol / Year: 2020Title: Light-coupled cryo-plunger for time-resolved cryo-EM. Authors: Nate Yoder / Farzad Jalali-Yazdi / Sigrid Noreng / Alexandra Houser / Isabelle Baconguis / Eric Gouaux /  Abstract: Proteins are dynamic molecules that can undergo rapid conformational rearrangements in response to stimuli. These structural changes are often critical to protein function, and thus elucidating time- ...Proteins are dynamic molecules that can undergo rapid conformational rearrangements in response to stimuli. These structural changes are often critical to protein function, and thus elucidating time-dependent conformational landscapes has been a long-standing goal of structural biology. To harness the power of single particle cryo-EM methods to enable 'time-resolved' structure determination, we have developed a light-coupled cryo-plunger that pairs flash-photolysis of caged ligands with rapid sample vitrification. The 'flash-plunger' consists of a high-power ultraviolet LED coupled with focusing optics and a motorized linear actuator, enabling the user to immobilize protein targets in vitreous ice within a programmable time window - as short as tens of milliseconds - after stimulus delivery. The flash-plunger is a simple, inexpensive and flexible tool to explore short-lived conformational states previously unobtainable by conventional sample preparation methods. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21385.map.gz emd_21385.map.gz | 168.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21385-v30.xml emd-21385-v30.xml emd-21385.xml emd-21385.xml | 12.5 KB 12.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21385.png emd_21385.png | 39.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21385 http://ftp.pdbj.org/pub/emdb/structures/EMD-21385 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21385 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21385 | HTTPS FTP |
-Validation report
| Summary document |  emd_21385_validation.pdf.gz emd_21385_validation.pdf.gz | 78.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21385_full_validation.pdf.gz emd_21385_full_validation.pdf.gz | 77.6 KB | Display | |
| Data in XML |  emd_21385_validation.xml.gz emd_21385_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21385 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21385 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21385 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21385 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21385.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21385.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map of an acid-sensing ion channel in a resting state at high pH. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
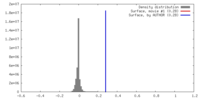
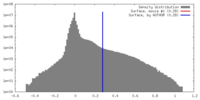
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Acid-sensing ion channel 1
| Entire | Name: Acid-sensing ion channel 1 |
|---|---|
| Components |
|
-Supramolecule #1: Acid-sensing ion channel 1
| Supramolecule | Name: Acid-sensing ion channel 1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 180 KDa |
-Macromolecule #1: Acid-sensing ion channel 1
| Macromolecule | Name: Acid-sensing ion channel 1 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MMDLKVDEEE VDSGQPVSIQ AFASSSTLHG ISHIFSYERL SLKRVVWALC FMGSLALLAL VCTNRIQYY FLYPHVTKLD EVAATRLTFP AVTFCNLNEF RFSRVTKNDL YHAGELLALL N NRYEIPDT QTADEKQLEI LQDKANFRNF KPKPFNMLEF YDRAGHDIRE ...String: MMDLKVDEEE VDSGQPVSIQ AFASSSTLHG ISHIFSYERL SLKRVVWALC FMGSLALLAL VCTNRIQYY FLYPHVTKLD EVAATRLTFP AVTFCNLNEF RFSRVTKNDL YHAGELLALL N NRYEIPDT QTADEKQLEI LQDKANFRNF KPKPFNMLEF YDRAGHDIRE MLLSCFFRGE QC SPEDFKV VFTRYGKCYT FNAGQDGKPR LITMKGGTGN GLEIMLDIQQ DEYLPVWGET DET SFEAGI KVQIHSQDEP PLIDQLGFGV APGFQTFVSC QEQRLIYLPP PWGDCKATTG DSEF YDTYS ITACRIDCET RYLVENCNCR MVHMPGDAPY CTPEQYKECA DPALDFLVEK DNEYC VCEM PCNVTRYGKE LSMVKIPSKA SAKYLAKKYN KSEQYIGENI LVLDIFFEAL NYETIE QKK AYEVAGLLGD IGGQMGLFIG ASILTVLELF DYAYEVIKHR LCRRGKCRKN HKRNNTD KG VALSMDDVKR HNPCESLRGH PAGMTYAANI LPHHPARGTF EDFTC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.00 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 70 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER Details: Specimen was frozen on a custom-made manual plunge apparatus in a cold room environment maintained at high humidity. Vitrification occurred following 25 ms of exposure to low power UV irradiation.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: Gctf |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2) / Number images used: 16673 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT / Software - Name: cryoSPARC (ver. 2) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 2) |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)