+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21141 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human CALHM2 | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of microglial cell activation / ATP export / calcium ion import / monoatomic cation channel activity / regulation of synaptic plasticity / positive regulation of apoptotic process / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.48 Å | |||||||||
 Authors Authors | Syrjanen JL / Chou TH / Furukawa H | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Structure and assembly of calcium homeostasis modulator proteins. Authors: Johanna L Syrjanen / Kevin Michalski / Tsung-Han Chou / Timothy Grant / Shanlin Rao / Noriko Simorowski / Stephen J Tucker / Nikolaus Grigorieff / Hiro Furukawa /   Abstract: The biological membranes of many cell types contain large-pore channels through which a wide variety of ions and metabolites permeate. Examples include connexin, innexin and pannexin, which form gap ...The biological membranes of many cell types contain large-pore channels through which a wide variety of ions and metabolites permeate. Examples include connexin, innexin and pannexin, which form gap junctions and/or bona fide cell surface channels. The most recently identified large-pore channels are the calcium homeostasis modulators (CALHMs), through which ions and ATP permeate in a voltage-dependent manner to control neuronal excitability, taste signaling and pathologies of depression and Alzheimer's disease. Despite such critical biological roles, the structures and patterns of their oligomeric assembly remain unclear. Here, we reveal the structures of two CALHMs, chicken CALHM1 and human CALHM2, by single-particle cryo-electron microscopy (cryo-EM), which show novel assembly of the four transmembrane helices into channels of octamers and undecamers, respectively. Furthermore, molecular dynamics simulations suggest that lipids can favorably assemble into a bilayer within the larger CALHM2 pore, but not within CALHM1, demonstrating the potential correlation between pore size, lipid accommodation and channel activity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21141.map.gz emd_21141.map.gz | 59.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21141-v30.xml emd-21141-v30.xml emd-21141.xml emd-21141.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21141_fsc.xml emd_21141_fsc.xml | 677 KB | Display |  FSC data file FSC data file |
| Images |  emd_21141.png emd_21141.png | 114 KB | ||
| Others |  emd_21141_additional.map.gz emd_21141_additional.map.gz | 58.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21141 http://ftp.pdbj.org/pub/emdb/structures/EMD-21141 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21141 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21141 | HTTPS FTP |
-Validation report
| Summary document |  emd_21141_validation.pdf.gz emd_21141_validation.pdf.gz | 538.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21141_full_validation.pdf.gz emd_21141_full_validation.pdf.gz | 538.3 KB | Display | |
| Data in XML |  emd_21141_validation.xml.gz emd_21141_validation.xml.gz | 117.7 KB | Display | |
| Data in CIF |  emd_21141_validation.cif.gz emd_21141_validation.cif.gz | 273.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21141 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21141 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21141 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21141 | HTTPS FTP |
-Related structure data
| Related structure data |  6vakMC  6vaiC  6valC  6vamC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_21141.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21141.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpened
| File | emd_21141_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
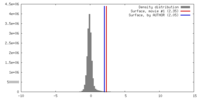
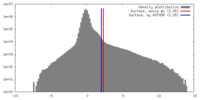
| Density Histograms |
- Sample components
Sample components
-Entire : Undecameric assembly of human calcium homeostasis modulator prote...
| Entire | Name: Undecameric assembly of human calcium homeostasis modulator protein 2 (CALHM2) in EDTA |
|---|---|
| Components |
|
-Supramolecule #1: Undecameric assembly of human calcium homeostasis modulator prote...
| Supramolecule | Name: Undecameric assembly of human calcium homeostasis modulator protein 2 (CALHM2) in EDTA type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
-Macromolecule #1: Calcium homeostasis modulator protein 2
| Macromolecule | Name: Calcium homeostasis modulator protein 2 / type: protein_or_peptide / ID: 1 / Number of copies: 11 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.468871 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAALIAENFR FLSLFFKSKD VMIFNGLVAL GTVGSQELFS VVAFHCPCSP ARNYLYGLAA IGVPALVLFI IGIILNNHTW NLVAECQHR RTKNCSAAPT FLLLSSILGR AAVAPVTWSV ISLLRGEAYV CALSEFVDPS SLTAREEHFP SAHATEILAR F PCKENPDN ...String: MAALIAENFR FLSLFFKSKD VMIFNGLVAL GTVGSQELFS VVAFHCPCSP ARNYLYGLAA IGVPALVLFI IGIILNNHTW NLVAECQHR RTKNCSAAPT FLLLSSILGR AAVAPVTWSV ISLLRGEAYV CALSEFVDPS SLTAREEHFP SAHATEILAR F PCKENPDN LSDFREEVSR RLRYESQLFG WLLIGVVAIL VFLTKCLKHY CSPLSYRQEA YWAQYRANED QLFQRTAEVH SR VLAANNV RRFFGFVALN KDDEELIANF PVEGTQPRPQ WNAITGVYLY RENQGLPLYS RLHKWAQGLA GNGAAPDNVE MAL LPSGSA WSHPQFEK |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 288.15 K / Instrument: FEI VITROBOT MARK IV / Details: Blot for 4 sec before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 70.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)