[English] 日本語
 Yorodumi
Yorodumi- EMDB-19552: cryo sub-tomogram average of Ald4 filaments from purified meiotic... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo sub-tomogram average of Ald4 filaments from purified meiotic yeast mitochondria | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | metabolic enzyme / filament / cryoET / mitochondria / CYTOSOLIC PROTEIN | |||||||||
| Biological species |   | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 44.0 Å | |||||||||
 Authors Authors | Hugener J / Xu J / Wettstein R / Ioannidi L / Velikov D / Wollweber F / Henggeler A / Matos J / Pilhofer M | |||||||||
| Funding support |  Switzerland, European Union, 2 items Switzerland, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: FilamentID reveals the composition and function of metabolic enzyme polymers during gametogenesis. Authors: Jannik Hugener / Jingwei Xu / Rahel Wettstein / Lydia Ioannidi / Daniel Velikov / Florian Wollweber / Adrian Henggeler / Joao Matos / Martin Pilhofer /   Abstract: Gamete formation and subsequent offspring development often involve extended phases of suspended cellular development or even dormancy. How cells adapt to recover and resume growth remains poorly ...Gamete formation and subsequent offspring development often involve extended phases of suspended cellular development or even dormancy. How cells adapt to recover and resume growth remains poorly understood. Here, we visualized budding yeast cells undergoing meiosis by cryo-electron tomography (cryoET) and discovered elaborate filamentous assemblies decorating the nucleus, cytoplasm, and mitochondria. To determine filament composition, we developed a "filament identification" (FilamentID) workflow that combines multiscale cryoET/cryo-electron microscopy (cryoEM) analyses of partially lysed cells or organelles. FilamentID identified the mitochondrial filaments as being composed of the conserved aldehyde dehydrogenase Ald4 and the nucleoplasmic/cytoplasmic filaments as consisting of acetyl-coenzyme A (CoA) synthetase Acs1. Structural characterization further revealed the mechanism underlying polymerization and enabled us to genetically perturb filament formation. Acs1 polymerization facilitates the recovery of chronologically aged spores and, more generally, the cell cycle re-entry of starved cells. FilamentID is broadly applicable to characterize filaments of unknown identity in diverse cellular contexts. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19552.map.gz emd_19552.map.gz | 119.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19552-v30.xml emd-19552-v30.xml emd-19552.xml emd-19552.xml | 13.5 KB 13.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_19552.png emd_19552.png | 13 KB | ||
| Filedesc metadata |  emd-19552.cif.gz emd-19552.cif.gz | 4.8 KB | ||
| Others |  emd_19552_half_map_1.map.gz emd_19552_half_map_1.map.gz emd_19552_half_map_2.map.gz emd_19552_half_map_2.map.gz | 106.3 KB 106.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19552 http://ftp.pdbj.org/pub/emdb/structures/EMD-19552 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19552 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19552 | HTTPS FTP |
-Validation report
| Summary document |  emd_19552_validation.pdf.gz emd_19552_validation.pdf.gz | 616.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19552_full_validation.pdf.gz emd_19552_full_validation.pdf.gz | 616.4 KB | Display | |
| Data in XML |  emd_19552_validation.xml.gz emd_19552_validation.xml.gz | 5.6 KB | Display | |
| Data in CIF |  emd_19552_validation.cif.gz emd_19552_validation.cif.gz | 6.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19552 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19552 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19552 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19552 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19552.map.gz / Format: CCP4 / Size: 128.9 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19552.map.gz / Format: CCP4 / Size: 128.9 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 11 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_19552_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19552_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ald4 polymer from purified meiotic yeast mitochondria
| Entire | Name: Ald4 polymer from purified meiotic yeast mitochondria |
|---|---|
| Components |
|
-Supramolecule #1: Ald4 polymer from purified meiotic yeast mitochondria
| Supramolecule | Name: Ald4 polymer from purified meiotic yeast mitochondria / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Aldehyde dehydrogenase 4
| Macromolecule | Name: Aldehyde dehydrogenase 4 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MFSRSTLCLK TSASSIGRL Q LRYFSHLP MT VPIKLPN GLE YEQPTG LFIN NKFVP SKQNK TFEV INPSTE EEI CHIYEGR ED DVEEAVQA A DRAFSNGSW NGIDPIDRGK ALYRLAELI E QDKDVIAS IE TLDNGKA ISS SRGDVD LVIN YLKSS ...String: MFSRSTLCLK TSASSIGRL Q LRYFSHLP MT VPIKLPN GLE YEQPTG LFIN NKFVP SKQNK TFEV INPSTE EEI CHIYEGR ED DVEEAVQA A DRAFSNGSW NGIDPIDRGK ALYRLAELI E QDKDVIAS IE TLDNGKA ISS SRGDVD LVIN YLKSS AGFAD KIDG RMIDTG RTH FSYTKRQ PL GVCGQIIP W NFPLLMWAW KIAPALVTGN TVVLKTAES T PLSALYVS KY IPQAGIP PGV INIVSG FGKI VGEAI TNHPK IKKV AFTGST ATG RHIYQSA AA GLKKVTLE L GGKSPNIVF ADAELKKAVQ NIILGIYYN S GEVCCAGS RV YVEESIY DKF IEEFKA ASES IKVGD PFDES TFQG AQTSQM QLN KILKYVD IG KNEGATLI T GGERLGSKG YFIKPTVFGD VKEDMRIVK E EIFGPVVT VT KFKSADE VIN MANDSE YGLA AGIHT SNINT ALKV ADRVNA GTV WINTYND FH HAVPFGGF N ASGLGREMS VDALQNYLQV KAVRAKLDE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 2.62 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 8.0 µm / Nominal defocus min: 8.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 44.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number subtomograms used: 564 |
|---|---|
| Extraction | Number tomograms: 3 / Number images used: 564 |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION |
 Movie
Movie Controller
Controller
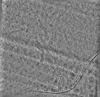










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)