+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1529 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Full-Length HIV-1 CA. | |||||||||
 Map data Map data | This is a map of full-length HIV-1 CA. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV-1 / CA / capsid / mature / retrovirus | |||||||||
| Function / homology |  Function and homology information Function and homology informationHIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding ...HIV-1 retropepsin / symbiont-mediated activation of host apoptosis / retroviral ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / viral penetration into host nucleus / host multivesicular body / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / host cell / viral nucleocapsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / symbiont-mediated suppression of host gene expression / viral translational frameshifting / symbiont entry into host cell / lipid binding / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / proteolysis / DNA binding / zinc ion binding Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | electron crystallography / cryo EM / negative staining / Resolution: 9.0 Å | |||||||||
 Authors Authors | Ganser-Pornillos BK / Cheng A / Yeager M | |||||||||
 Citation Citation |  Journal: Cell / Year: 2007 Journal: Cell / Year: 2007Title: Structure of full-length HIV-1 CA: a model for the mature capsid lattice. Authors: Barbie K Ganser-Pornillos / Anchi Cheng / Mark Yeager /  Abstract: The capsids of mature retroviruses perform the essential function of organizing the viral genome for efficient replication. These capsids are modeled as fullerene structures composed of closed ...The capsids of mature retroviruses perform the essential function of organizing the viral genome for efficient replication. These capsids are modeled as fullerene structures composed of closed hexameric arrays of the viral CA protein, but a high-resolution structure of the lattice has remained elusive. A three-dimensional map of two-dimensional crystals of the R18L mutant of HIV-1 CA was derived by electron cryocrystallography. The docking of high-resolution domain structures into the map yielded the first unambiguous model for full-length HIV-1 CA. Three important protein-protein assembly interfaces are required for capsid formation. Each CA hexamer is composed of an inner ring of six N-terminal domains and an outer ring of C-terminal domains that form dimeric linkers connecting neighboring hexamers. Interactions between the two domains of CA further stabilize the hexamer and provide a structural explanation for the mechanism of action of known HIV-1 assembly inhibitors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1529.map.gz emd_1529.map.gz | 5.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1529-v30.xml emd-1529-v30.xml emd-1529.xml emd-1529.xml | 11.7 KB 11.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1529.gif 1529.gif | 69.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1529 http://ftp.pdbj.org/pub/emdb/structures/EMD-1529 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1529 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1529 | HTTPS FTP |
-Related structure data
| Related structure data |  3dikMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1529.map.gz / Format: CCP4 / Size: 14.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1529.map.gz / Format: CCP4 / Size: 14.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
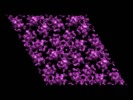
| Annotation | This is a map of full-length HIV-1 CA. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X: 1.03 Å / Y: 1.03 Å / Z: 0.92437 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 168 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : HIV-1 CA R18L
| Entire | Name: HIV-1 CA R18L |
|---|---|
| Components |
|
-Supramolecule #1000: HIV-1 CA R18L
| Supramolecule | Name: HIV-1 CA R18L / type: sample / ID: 1000 / Oligomeric state: 6 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 25 KDa / Theoretical: 25 KDa / Method: mass spectrometry of monomer |
-Macromolecule #1: HIV-1 CA protein
| Macromolecule | Name: HIV-1 CA protein / type: protein_or_peptide / ID: 1 / Name.synonym: HIV-1 CA protein / Details: mass spectrometry of virus / Oligomeric state: hexamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 / synonym: HIV-1 Human immunodeficiency virus 1 / synonym: HIV-1 |
| Molecular weight | Experimental: 25 KDa / Theoretical: 25 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | electron crystallography |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Concentration | 32 mg/mL |
|---|---|
| Buffer | pH: 6.5 Details: 17.5% (w/v) PEG 20,000, 50mM sodium cacodylate, 100mM Calcium acetate |
| Staining | Type: NEGATIVE Details: Grids were absorbed with crystals, washed with 0.1M KCl, blotted, and flash frozen in liquid ethane |
| Grid | Details: 300 mesh molybdemun |
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER / Method: blot for 2 sec before plungin |
| Details | crstal grown by direct addition of PEG solution |
| Crystal formation | Details: crstal grown by direct addition of PEG solution |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 100,000 times magnification |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 43 / Average electron dose: 15 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Gatan 626 / Specimen holder model: GATAN LIQUID NITROGEN / Tilt angle max: 40 / Tilt series - Axis1 - Min angle: 0 ° / Tilt series - Axis1 - Max angle: 40 ° |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 9.0 Å / Software - Name: MRC |
|---|---|
| Crystal parameters | Unit cell - A: 92.7 Å / Unit cell - B: 92.7 Å / Unit cell - C: 110 Å / Unit cell - γ: 120 ° / Unit cell - α: 90 ° / Unit cell - β: 90 ° / Plane group: P 6 |
| CTF correction | Details: each image |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: Situs |
| Details | Protocol: Rigid Body |
| Refinement | Space: RECIPROCAL / Protocol: RIGID BODY FIT |
| Output model |  PDB-3dik: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)























