+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PucA-LH2 complex from Rps. palustris | |||||||||
 Map data Map data | LH2 from Rps. palustris. | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationorganelle inner membrane / plasma membrane light-harvesting complex / bacteriochlorophyll binding / photosynthesis, light reaction / electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity / membrane => GO:0016020 / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Rhodopseudomonas palustris ATCC 17001 (phototrophic) Rhodopseudomonas palustris ATCC 17001 (phototrophic) | |||||||||
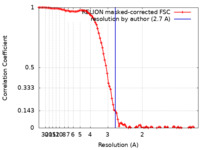
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Qian P / Cogdell RJ | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Cryo-EM structures of light-harvesting 2 complexes from reveal the molecular origin of absorption tuning. Authors: Pu Qian / Cam T Nguyen-Phan / Alastair T Gardiner / Tristan I Croll / Aleksander W Roszak / June Southall / Philip J Jackson / Cvetelin Vasilev / Pablo Castro-Hartmann / Kasim Sader / C Neil ...Authors: Pu Qian / Cam T Nguyen-Phan / Alastair T Gardiner / Tristan I Croll / Aleksander W Roszak / June Southall / Philip J Jackson / Cvetelin Vasilev / Pablo Castro-Hartmann / Kasim Sader / C Neil Hunter / Richard J Cogdell /   Abstract: The genomes of some purple photosynthetic bacteria contain a multigene family encoding a series of α- and β-polypeptides that together form a heterogeneous antenna of light-harvesting 2 (LH2) ...The genomes of some purple photosynthetic bacteria contain a multigene family encoding a series of α- and β-polypeptides that together form a heterogeneous antenna of light-harvesting 2 (LH2) complexes. To unravel this complexity, we generated four sets of deletion mutants in , each encoding a single type of gene pair and enabling the purification of complexes designated as PucA-LH2, PucB-LH2, PucD-LH2, and PucE-LH2. The structures of all four purified LH2 complexes were determined by cryogenic electron microscopy (cryo-EM) at resolutions ranging from 2.7 to 3.6 Å. Uniquely, each of these complexes contains a hitherto unknown polypeptide, γ, that forms an extended undulating ribbon that lies in the plane of the membrane and that encloses six of the nine LH2 αβ-subunits. The γ-subunit, which is located near to the cytoplasmic side of the complex, breaks the C9 symmetry of the LH2 complex and binds six extra bacteriochlorophylls (BChls) that enhance the 800-nm absorption of each complex. The structures show that all four complexes have two complete rings of BChls, conferring absorption bands centered at 800 and 850 nm on the PucA-LH2, PucB-LH2, and PucE-LH2 complexes, but, unusually, the PucD-LH2 antenna has only a single strong near-infared (NIR) absorption peak at 803 nm. Comparison of the cryo-EM structures of these LH2 complexes reveals altered patterns of hydrogen bonds between LH2 αβ-side chains and the bacteriochlorin rings, further emphasizing the major role that H bonds play in spectral tuning of bacterial antenna complexes. #1:  Journal: Acta Crystallogr., Sect. D: Biol. Crystallogr. / Year: 2018 Journal: Acta Crystallogr., Sect. D: Biol. Crystallogr. / Year: 2018Title: Real-space refinement in PHENIX for cryo-EM and crystallography Authors: Qian P / Cogdell RJ #2:  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structures of light harvesting complex 2 from Rhodopseudomonas palustris: a molecular origin of spectroscopic variation Authors: Qian P / Cogdell R | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14633.map.gz emd_14633.map.gz | 9.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14633-v30.xml emd-14633-v30.xml emd-14633.xml emd-14633.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14633_fsc.xml emd_14633_fsc.xml | 9.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_14633.png emd_14633.png | 81.3 KB | ||
| Others |  emd_14633_half_map_1.map.gz emd_14633_half_map_1.map.gz emd_14633_half_map_2.map.gz emd_14633_half_map_2.map.gz | 58.5 MB 58.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14633 http://ftp.pdbj.org/pub/emdb/structures/EMD-14633 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14633 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14633 | HTTPS FTP |
-Validation report
| Summary document |  emd_14633_validation.pdf.gz emd_14633_validation.pdf.gz | 714.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14633_full_validation.pdf.gz emd_14633_full_validation.pdf.gz | 714.5 KB | Display | |
| Data in XML |  emd_14633_validation.xml.gz emd_14633_validation.xml.gz | 16.7 KB | Display | |
| Data in CIF |  emd_14633_validation.cif.gz emd_14633_validation.cif.gz | 21.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14633 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14633 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14633 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14633 | HTTPS FTP |
-Related structure data
| Related structure data |  7zcuMC  7zdiC  7ze3C  7ze8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14633.map.gz / Format: CCP4 / Size: 75.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14633.map.gz / Format: CCP4 / Size: 75.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LH2 from Rps. palustris. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.65 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: The first half map, z-flipped.
| File | emd_14633_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The first half map, z-flipped. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: The second half map, z-flipped.
| File | emd_14633_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The second half map, z-flipped. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PucA-LH2 from Rps. palustris
| Entire | Name: PucA-LH2 from Rps. palustris |
|---|---|
| Components |
|
-Supramolecule #1: PucA-LH2 from Rps. palustris
| Supramolecule | Name: PucA-LH2 from Rps. palustris / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / Details: Delt-pucBA(bced) |
|---|---|
| Source (natural) | Organism:  Rhodopseudomonas palustris ATCC 17001 (phototrophic) Rhodopseudomonas palustris ATCC 17001 (phototrophic) |
-Macromolecule #1: Light-harvesting protein B-800-850 alpha chain
| Macromolecule | Name: Light-harvesting protein B-800-850 alpha chain / type: protein_or_peptide / ID: 1 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rhodopseudomonas palustris ATCC 17001 (phototrophic) Rhodopseudomonas palustris ATCC 17001 (phototrophic)Strain: ATCC 17001 |
| Molecular weight | Theoretical: 6.443515 KDa |
| Sequence | String: (CXM)NQARIWTVV KPTVGLPLLL GSVTVIAILV HFAVLSHTTW FSKYWNGKAA AIESSVNVG |
-Macromolecule #2: Light-harvesting protein B-800-850 beta chain
| Macromolecule | Name: Light-harvesting protein B-800-850 beta chain / type: protein_or_peptide / ID: 2 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rhodopseudomonas palustris ATCC 17001 (phototrophic) Rhodopseudomonas palustris ATCC 17001 (phototrophic)Strain: ATCC 17001 |
| Molecular weight | Theoretical: 5.288061 KDa |
| Sequence | String: MADKTLTGLT VEESEELHKH VIDGTRIFGA IAIVAHFLAY VYSPWLH |
-Macromolecule #3: PucA-LH2-gamma
| Macromolecule | Name: PucA-LH2-gamma / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rhodopseudomonas palustris ATCC 17001 (phototrophic) Rhodopseudomonas palustris ATCC 17001 (phototrophic)Strain: ATCC 17001 |
| Molecular weight | Theoretical: 10.848179 KDa |
| Sequence | String: MSEEYKGHSG HPLILKQEGE YKGYSGEPLI LKQEGEYKGY SGTPLILEQK GEYQSFSGTP LILKQEGEYR GFSGAPLILK QDGEYKSFS GYPLLLNI |
-Macromolecule #4: BACTERIOCHLOROPHYLL A
| Macromolecule | Name: BACTERIOCHLOROPHYLL A / type: ligand / ID: 4 / Number of copies: 33 / Formula: BCL |
|---|---|
| Molecular weight | Theoretical: 911.504 Da |
| Chemical component information |  ChemComp-BCL: |
-Macromolecule #5: 1,2-Dihydro-psi,psi-caroten-1-ol
| Macromolecule | Name: 1,2-Dihydro-psi,psi-caroten-1-ol / type: ligand / ID: 5 / Number of copies: 9 / Formula: IRM |
|---|---|
| Molecular weight | Theoretical: 554.888 Da |
| Chemical component information |  ChemComp-IRM: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 9 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.0 mg/mL |
|---|---|
| Buffer | pH: 8 / Component - Concentration: 20.0 mM / Component - Formula: C4H11NO3 / Component - Name: Tris.HCl / Details: 0.1% LDAO in 20 mM Tris.Cl pH 8.0 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Chamber humidity: 100%; Chamber temperature: 4 oC; Blotting time: 2.5; Blotting force: 3; Wait time: 30 sec. |
| Details | protein was purified using LDAO detergent. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 80.0 K |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 4865 / Average exposure time: 1.4 sec. / Average electron dose: 42.03 e/Å2 / Details: Images were collected in AFIS mode. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)