+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13088 | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
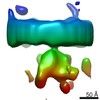
| Title | Mature HIV-1 matrix structure | |||||||||||||||||||||||||||
 Map data Map data | Mature HIV-1 MA structure | |||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationviral process / viral nucleocapsid / host cell cytoplasm / host cell nucleus / structural molecule activity / virion membrane / RNA binding / zinc ion binding / ATP binding / cytoplasm Similarity search - Function | |||||||||||||||||||||||||||
| Biological species |   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||||||||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 7.0 Å | |||||||||||||||||||||||||||
 Authors Authors | Qu K / Ke ZL / Zila V / Anders-Oesswein M / Glass B / Muecksch F / Mueller R / Schultz C / Mueller B / Kraeusslich HG / Briggs JAG | |||||||||||||||||||||||||||
| Funding support | European Union,  United Kingdom, United Kingdom,  Germany, Germany,  United States, 8 items United States, 8 items
| |||||||||||||||||||||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Maturation of the matrix and viral membrane of HIV-1. Authors: Kun Qu / Zunlong Ke / Vojtech Zila / Maria Anders-Össwein / Bärbel Glass / Frauke Mücksch / Rainer Müller / Carsten Schultz / Barbara Müller / Hans-Georg Kräusslich / John A G Briggs /    Abstract: Gag, the primary structural protein of HIV-1, is recruited to the plasma membrane for virus assembly by its matrix (MA) domain. Gag is subsequently cleaved into its component domains, causing ...Gag, the primary structural protein of HIV-1, is recruited to the plasma membrane for virus assembly by its matrix (MA) domain. Gag is subsequently cleaved into its component domains, causing structural maturation to repurpose the virion for cell entry. We determined the structure and arrangement of MA within immature and mature HIV-1 through cryo-electron tomography. We found that MA rearranges between two different hexameric lattices upon maturation. In mature HIV-1, a lipid extends out of the membrane to bind with a pocket in MA. Our data suggest that proteolytic maturation of HIV-1 not only assembles the viral capsid surrounding the genome but also repurposes the membrane-bound MA lattice for an entry or postentry function and results in the partial removal of up to 2500 lipids from the viral membrane. | |||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13088.map.gz emd_13088.map.gz | 17.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13088-v30.xml emd-13088-v30.xml emd-13088.xml emd-13088.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13088.png emd_13088.png | 278.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13088 http://ftp.pdbj.org/pub/emdb/structures/EMD-13088 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13088 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13088 | HTTPS FTP |
-Related structure data
| Related structure data |  7ovrMC  7ovqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13088.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13088.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mature HIV-1 MA structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human immunodeficiency virus 1
| Entire | Name:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
|---|---|
| Components |
|
-Supramolecule #1: Human immunodeficiency virus 1
| Supramolecule | Name: Human immunodeficiency virus 1 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: cHIV MA-SP1 Gag proteolytic cleavage mutant virus particles purified from HEK293T cells. NCBI-ID: 11676 / Sci species name: Human immunodeficiency virus 1 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Host system | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: HIV-1 matrix
| Macromolecule | Name: HIV-1 matrix / type: protein_or_peptide / ID: 1 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 13.084983 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GARASVLSGG ELDKWEKIRL RPGGKKQYKL KHIVWASREL ERFAVNPGLL ETSEGCRQIL GQLQPSLQTG SEELRSLYNT IAVLYCVHQ RIDVKDTKEA LDKIEEEQNK SKKKAQ |
-Macromolecule #2: MYRISTIC ACID
| Macromolecule | Name: MYRISTIC ACID / type: ligand / ID: 2 / Number of copies: 24 / Formula: MYR |
|---|---|
| Molecular weight | Theoretical: 228.371 Da |
| Chemical component information |  ChemComp-MYR: |
-Macromolecule #3: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(o...
| Macromolecule | Name: [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate type: ligand / ID: 3 / Number of copies: 24 / Formula: PIO |
|---|---|
| Molecular weight | Theoretical: 746.566 Da |
| Chemical component information |  ChemComp-PIO: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
| Details | cHIV MA-SP1 Gag proteolytic cleavage mutant virus particles purified from HEK293T cells. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 3.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 7.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: AV3 / Number subtomograms used: 19586 |
|---|---|
| Extraction | Number tomograms: 65 / Number images used: 119343 / Software - Name:  MATLAB MATLAB |
| CTF correction | Software: (Name: CTFFIND, NOVACTF) |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller