+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12928 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
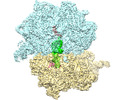
| Title | CspA-27 cotranslational folding intermediate 2 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | cotranslational folding / CspA27-2 / RIBOSOME | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationtranscriptional attenuation / endoribonuclease inhibitor activity / RNA-binding transcription regulator activity / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / assembly of large subunit precursor of preribosome / ribosome assembly / cytosolic ribosome assembly ...transcriptional attenuation / endoribonuclease inhibitor activity / RNA-binding transcription regulator activity / negative regulation of cytoplasmic translation / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / assembly of large subunit precursor of preribosome / ribosome assembly / cytosolic ribosome assembly / DNA-templated transcription termination / response to radiation / mRNA 5'-UTR binding / large ribosomal subunit / transferase activity / ribosomal small subunit assembly / small ribosomal subunit / 5S rRNA binding / ribosomal large subunit assembly / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / response to antibiotic / negative regulation of DNA-templated transcription / mRNA binding / DNA binding / RNA binding / zinc ion binding / metal ion binding / membrane / cytosol / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | ||||||||||||
 Authors Authors | Agirrezabala X / Samatova E | ||||||||||||
| Funding support |  Spain, European Union, 3 items Spain, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: EMBO J / Year: 2022 Journal: EMBO J / Year: 2022Title: A switch from α-helical to β-strand conformation during co-translational protein folding. Authors: Xabier Agirrezabala / Ekaterina Samatova / Meline Macher / Marija Liutkute / Manisankar Maiti / David Gil-Carton / Jiri Novacek / Mikel Valle / Marina V Rodnina /    Abstract: Cellular proteins begin to fold as they emerge from the ribosome. The folding landscape of nascent chains is not only shaped by their amino acid sequence but also by the interactions with the ...Cellular proteins begin to fold as they emerge from the ribosome. The folding landscape of nascent chains is not only shaped by their amino acid sequence but also by the interactions with the ribosome. Here, we combine biophysical methods with cryo-EM structure determination to show that folding of a β-barrel protein begins with formation of a dynamic α-helix inside the ribosome. As the growing peptide reaches the end of the tunnel, the N-terminal part of the nascent chain refolds to a β-hairpin structure that remains dynamic until its release from the ribosome. Contacts with the ribosome and structure of the peptidyl transferase center depend on nascent chain conformation. These results indicate that proteins may start out as α-helices inside the tunnel and switch into their native folds only as they emerge from the ribosome. Moreover, the correlation of nascent chain conformations with reorientation of key residues of the ribosomal peptidyl-transferase center suggest that protein folding could modulate ribosome activity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12928.map.gz emd_12928.map.gz | 193.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12928-v30.xml emd-12928-v30.xml emd-12928.xml emd-12928.xml | 87.1 KB 87.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12928.png emd_12928.png | 176.6 KB | ||
| Masks |  emd_12928_msk_1.map emd_12928_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12928.cif.gz emd-12928.cif.gz | 15.8 KB | ||
| Others |  emd_12928_half_map_1.map.gz emd_12928_half_map_1.map.gz emd_12928_half_map_2.map.gz emd_12928_half_map_2.map.gz | 194.9 MB 194.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12928 http://ftp.pdbj.org/pub/emdb/structures/EMD-12928 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12928 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12928 | HTTPS FTP |
-Validation report
| Summary document |  emd_12928_validation.pdf.gz emd_12928_validation.pdf.gz | 164.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12928_full_validation.pdf.gz emd_12928_full_validation.pdf.gz | 164.5 KB | Display | |
| Data in XML |  emd_12928_validation.xml.gz emd_12928_validation.xml.gz | 571 B | Display | |
| Data in CIF |  emd_12928_validation.cif.gz emd_12928_validation.cif.gz | 483 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12928 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12928 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12928 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12928 | HTTPS FTP |
-Related structure data
| Related structure data |  7oifMC  7nwwC  7oigC  7oiiC  7ot5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12928.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12928.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
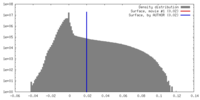
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12928_msk_1.map emd_12928_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
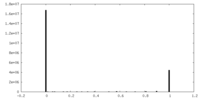
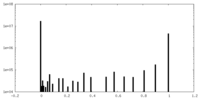
| Density Histograms |
-Half map: #2
| File | emd_12928_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
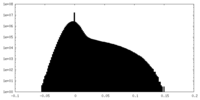
| Density Histograms |
-Half map: #1
| File | emd_12928_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
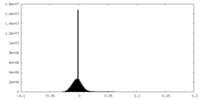
| Density Histograms |
- Sample components
Sample components
+Entire : 70S-CspA27-2
+Supramolecule #1: 70S-CspA27-2
+Macromolecule #1: 23S rRNA
+Macromolecule #2: 16S rRNA
+Macromolecule #3: 5S rRNA
+Macromolecule #4: mRNA
+Macromolecule #54: tRNA-Ser
+Macromolecule #5: 50S ribosomal protein L2
+Macromolecule #6: 50S ribosomal protein L3
+Macromolecule #7: 50S ribosomal protein L4
+Macromolecule #8: 50S ribosomal protein L5
+Macromolecule #9: 50S ribosomal protein L6
+Macromolecule #10: 50S ribosomal protein L9
+Macromolecule #11: 50S ribosomal protein L13
+Macromolecule #12: 50S ribosomal protein L14
+Macromolecule #13: 50S ribosomal protein L15
+Macromolecule #14: 50S ribosomal protein L16
+Macromolecule #15: 50S ribosomal protein L17
+Macromolecule #16: 50S ribosomal protein L18
+Macromolecule #17: 50S ribosomal protein L19
+Macromolecule #18: 50S ribosomal protein L20
+Macromolecule #19: 50S ribosomal protein L21
+Macromolecule #20: 50S ribosomal protein L22
+Macromolecule #21: 50S ribosomal protein L23
+Macromolecule #22: 50S ribosomal protein L24
+Macromolecule #23: 50S ribosomal protein L25
+Macromolecule #24: 50S ribosomal protein L27
+Macromolecule #25: 50S ribosomal protein L28
+Macromolecule #26: 50S ribosomal protein L29
+Macromolecule #27: 50S ribosomal protein L30
+Macromolecule #28: 50S ribosomal protein L31
+Macromolecule #29: 50S ribosomal protein L32
+Macromolecule #30: 50S ribosomal protein L33
+Macromolecule #31: 50S ribosomal protein L34
+Macromolecule #32: 50S ribosomal protein L35
+Macromolecule #33: 50S ribosomal protein L36
+Macromolecule #34: 30S ribosomal protein S2
+Macromolecule #35: 30S ribosomal protein S3
+Macromolecule #36: 30S ribosomal protein S4
+Macromolecule #37: 30S ribosomal protein S5
+Macromolecule #38: 30S ribosomal protein S6
+Macromolecule #39: 30S ribosomal protein S7
+Macromolecule #40: 30S ribosomal protein S8
+Macromolecule #41: 30S ribosomal protein S9
+Macromolecule #42: 30S ribosomal protein S10
+Macromolecule #43: 30S ribosomal protein S11
+Macromolecule #44: 30S ribosomal protein S12
+Macromolecule #45: 30S ribosomal protein S13
+Macromolecule #46: 30S ribosomal protein S14
+Macromolecule #47: 30S ribosomal protein S15
+Macromolecule #48: 30S ribosomal protein S16
+Macromolecule #49: 30S ribosomal protein S17
+Macromolecule #50: 30S ribosomal protein S18
+Macromolecule #51: 30S ribosomal protein S19
+Macromolecule #52: 30S ribosomal protein S20
+Macromolecule #53: 30S ribosomal protein S21
+Macromolecule #55: CspA transcriptional activator
+Macromolecule #56: MAGNESIUM ION
+Macromolecule #57: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 50 mM Tris-HCl, 70 mM NH4Cl, 30 mM KCl, 3.5 mM MgCl2, 8 mM putrescine, 0.5 mM spermidine, 1 mM DTT, 1 mM GTP, pH 7.5 |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 99 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 2.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | The software LocScale for model-based local density sharpening was used as implemented in the CCP-EM suite, version 1.3.0 |
| Refinement | Space: REAL / Protocol: OTHER |
| Output model |  PDB-7oif: |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)