+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12570 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | AL amyloid fibril from a lambda 1 light chain | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | amyloid / antibody / systemic amyloidosis / light chain / IMMUNE SYSTEM | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Karimi Farsijani S / Radamaker L | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Role of mutations and post-translational modifications in systemic AL amyloidosis studied by cryo-EM. Authors: Lynn Radamaker / Sara Karimi-Farsijani / Giada Andreotti / Julian Baur / Matthias Neumann / Sarah Schreiner / Natalie Berghaus / Raoul Motika / Christian Haupt / Paul Walther / Volker ...Authors: Lynn Radamaker / Sara Karimi-Farsijani / Giada Andreotti / Julian Baur / Matthias Neumann / Sarah Schreiner / Natalie Berghaus / Raoul Motika / Christian Haupt / Paul Walther / Volker Schmidt / Stefanie Huhn / Ute Hegenbart / Stefan O Schönland / Sebastian Wiese / Clarissa Read / Matthias Schmidt / Marcus Fändrich /  Abstract: Systemic AL amyloidosis is a rare disease that is caused by the misfolding of immunoglobulin light chains (LCs). Potential drivers of amyloid formation in this disease are post-translational ...Systemic AL amyloidosis is a rare disease that is caused by the misfolding of immunoglobulin light chains (LCs). Potential drivers of amyloid formation in this disease are post-translational modifications (PTMs) and the mutational changes that are inserted into the LCs by somatic hypermutation. Here we present the cryo electron microscopy (cryo-EM) structure of an ex vivo λ1-AL amyloid fibril whose deposits disrupt the ordered cardiomyocyte structure in the heart. The fibril protein contains six mutational changes compared to the germ line and three PTMs (disulfide bond, N-glycosylation and pyroglutamylation). Our data imply that the disulfide bond, glycosylation and mutational changes contribute to determining the fibril protein fold and help to generate a fibril morphology that is able to withstand proteolytic degradation inside the body. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12570.map.gz emd_12570.map.gz | 58.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12570-v30.xml emd-12570-v30.xml emd-12570.xml emd-12570.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
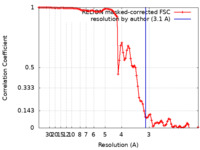
| FSC (resolution estimation) |  emd_12570_fsc.xml emd_12570_fsc.xml | 9.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_12570.png emd_12570.png | 108.7 KB | ||
| Filedesc metadata |  emd-12570.cif.gz emd-12570.cif.gz | 6 KB | ||
| Others |  emd_12570_half_map_1.map.gz emd_12570_half_map_1.map.gz emd_12570_half_map_2.map.gz emd_12570_half_map_2.map.gz | 52 MB 51.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12570 http://ftp.pdbj.org/pub/emdb/structures/EMD-12570 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12570 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12570 | HTTPS FTP |
-Related structure data
| Related structure data |  7nslMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10730 (Title: AL amyloid fibril from a glycosylated lambda 1 light chain EMPIAR-10730 (Title: AL amyloid fibril from a glycosylated lambda 1 light chainData size: 495.9 Data #1: Unaligned multi-frame micrographs of AL amyloidosis extracted ex vivo amyloid fibrils patient FOR001 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12570.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12570.map.gz / Format: CCP4 / Size: 67 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #2
| File | emd_12570_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
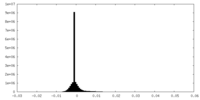
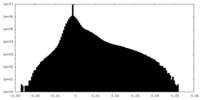
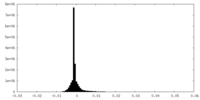
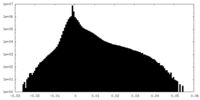
| Density Histograms |
-Half map: #1
| File | emd_12570_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Amyloid fibril of an antibody lambda 1 immunoglobulin light chain
| Entire | Name: Amyloid fibril of an antibody lambda 1 immunoglobulin light chain |
|---|---|
| Components |
|
-Supramolecule #1: Amyloid fibril of an antibody lambda 1 immunoglobulin light chain
| Supramolecule | Name: Amyloid fibril of an antibody lambda 1 immunoglobulin light chain type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Extracted fibrils from the explanted heart of a patient suffering from systemic AL amyloidosis |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Heart / Tissue: Heart muscle Homo sapiens (human) / Organ: Heart / Tissue: Heart muscle |
-Macromolecule #1: Amyloid lambda1 light chain
| Macromolecule | Name: Amyloid lambda1 light chain / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Organ: Heart / Tissue: Heart muscle tissue Homo sapiens (human) / Organ: Heart / Tissue: Heart muscle tissue |
| Molecular weight | Theoretical: 12.122354 KDa |
| Sequence | String: QSVLTQPPSV SAAPGQNVTI SCSGSSSNIG NNYVSWYQQL PGTAPKLLIY ETDKRPSGIP DRFSGSKSGT SATLGITGLQ TADEADYYC GTWESSLLAG VFGGGTKLTV LGQPKAAPS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Component - Formula: H2O / Component - Name: Distilled water |
|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK III / Details: blot for 9s before plunging. |
| Details | Sample in pure water, pH not determined |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Number grids imaged: 1 / Number real images: 3032 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Secondary structure restraints and NCS were applied during refinement |
|---|---|
| Refinement | Space: REAL / Protocol: OTHER Target criteria: REAL-SPACE (WEIGHTED MAP SUM AT ATOM CENTERS) |
| Output model |  PDB-7nsl: |
 Movie
Movie Controller
Controller














 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)