[English] 日本語
 Yorodumi
Yorodumi- EMDB-12286: Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Gu... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12286 | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
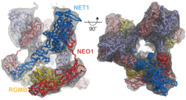
| Title | Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Guidance Molecule B complex | |||||||||||||||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||||||||||||||
 Keywords Keywords | Signal transduction / cell surface receptors / neuron regeneration / cell migration / Netrin / Neogenin / Repulsive Guidance Molecule / complex structure / protein-protein interactions / SIGNALING PROTEIN | |||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of glial cell migration / DSCAM interactions / chemorepulsion of axon / Cdc42 protein signal transduction / anterior/posterior axon guidance / Netrin-1 signaling / motor neuron migration / Regulation of commissural axon pathfinding by SLIT and ROBO / Role of second messengers in netrin-1 signaling / negative regulation of axon extension ...regulation of glial cell migration / DSCAM interactions / chemorepulsion of axon / Cdc42 protein signal transduction / anterior/posterior axon guidance / Netrin-1 signaling / motor neuron migration / Regulation of commissural axon pathfinding by SLIT and ROBO / Role of second messengers in netrin-1 signaling / negative regulation of axon extension / mammary gland duct morphogenesis / substrate-dependent cell migration, cell extension / Netrin mediated repulsion signals / DCC mediated attractive signaling / positive regulation of cell motility / inner ear morphogenesis / nuclear migration / regulation of synapse assembly / endoplasmic reticulum-Golgi intermediate compartment / basement membrane / positive regulation of glial cell proliferation / BMP signaling pathway / positive regulation of axon extension / glial cell proliferation / side of membrane / coreceptor activity / cell-cell adhesion / actin cytoskeleton / Ras protein signal transduction / DNA-binding transcription factor activity, RNA polymerase II-specific / cell adhesion / RNA polymerase II cis-regulatory region sequence-specific DNA binding / membrane raft / apoptotic process / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / glutamatergic synapse / signal transduction / extracellular region / nucleoplasm / identical protein binding / plasma membrane / cytosol Similarity search - Function | |||||||||||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||||||||||||||||||||
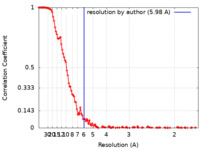
| Method | single particle reconstruction / cryo EM / Resolution: 5.98 Å | |||||||||||||||||||||||||||||||||
 Authors Authors | Robinson RA / Griffiths SC | |||||||||||||||||||||||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Netherlands, 10 items Netherlands, 10 items
| |||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2021 Journal: Cell / Year: 2021Title: Simultaneous binding of Guidance Cues NET1 and RGM blocks extracellular NEO1 signaling. Authors: Ross A Robinson / Samuel C Griffiths / Lieke L van de Haar / Tomas Malinauskas / Eljo Y van Battum / Pavol Zelina / Rebekka A Schwab / Dimple Karia / Lina Malinauskaite / Sara Brignani / ...Authors: Ross A Robinson / Samuel C Griffiths / Lieke L van de Haar / Tomas Malinauskas / Eljo Y van Battum / Pavol Zelina / Rebekka A Schwab / Dimple Karia / Lina Malinauskaite / Sara Brignani / Marleen H van den Munkhof / Özge Düdükcü / Anna A De Ruiter / Dianne M A Van den Heuvel / Benjamin Bishop / Jonathan Elegheert / A Radu Aricescu / R Jeroen Pasterkamp / Christian Siebold /   Abstract: During cell migration or differentiation, cell surface receptors are simultaneously exposed to different ligands. However, it is often unclear how these extracellular signals are integrated. Neogenin ...During cell migration or differentiation, cell surface receptors are simultaneously exposed to different ligands. However, it is often unclear how these extracellular signals are integrated. Neogenin (NEO1) acts as an attractive guidance receptor when the Netrin-1 (NET1) ligand binds, but it mediates repulsion via repulsive guidance molecule (RGM) ligands. Here, we show that signal integration occurs through the formation of a ternary NEO1-NET1-RGM complex, which triggers reciprocal silencing of downstream signaling. Our NEO1-NET1-RGM structures reveal a "trimer-of-trimers" super-assembly, which exists in the cell membrane. Super-assembly formation results in inhibition of RGMA-NEO1-mediated growth cone collapse and RGMA- or NET1-NEO1-mediated neuron migration, by preventing formation of signaling-compatible RGM-NEO1 complexes and NET1-induced NEO1 ectodomain clustering. These results illustrate how simultaneous binding of ligands with opposing functions, to a single receptor, does not lead to competition for binding, but to formation of a super-complex that diminishes their functional outputs. | |||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12286.map.gz emd_12286.map.gz | 24.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12286-v30.xml emd-12286-v30.xml emd-12286.xml emd-12286.xml | 33.6 KB 33.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12286_fsc.xml emd_12286_fsc.xml | 16.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_12286.png emd_12286.png | 461 KB | ||
| Masks |  emd_12286_msk_1.map emd_12286_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12286.cif.gz emd-12286.cif.gz | 9 KB | ||
| Others |  emd_12286_additional_1.map.gz emd_12286_additional_1.map.gz emd_12286_half_map_1.map.gz emd_12286_half_map_1.map.gz emd_12286_half_map_2.map.gz emd_12286_half_map_2.map.gz | 155.4 MB 131.8 MB 131.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12286 http://ftp.pdbj.org/pub/emdb/structures/EMD-12286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12286 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12286 | HTTPS FTP |
-Related structure data
| Related structure data |  7ndgMC  7ne0C  7ne1C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10637 (Title: Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Guidance Molecule B complex EMPIAR-10637 (Title: Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Guidance Molecule B complexData size: 1.2 TB Data #1: Unaligned multi-frame micrographs of the ternary Netrin 1-Neogenin 1-Repulsive Guidance Molecule B complex [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12286.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12286.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12286_msk_1.map emd_12286_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
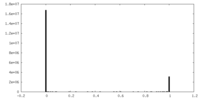
| Density Histograms |
-Additional map: #1
| File | emd_12286_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_12286_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_12286_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Gu...
+Supramolecule #1: Cryo-EM structure of the ternary Netrin 1-Neogenin 1-Repulsive Gu...
+Supramolecule #2: Netrin 1
+Supramolecule #3: Neogenin
+Supramolecule #4: Repulsive Guidance Molecule B
+Supramolecule #5: Repulsive Guidance Molecule B C-terminal region (chain D)
+Macromolecule #1: Netrin-1
+Macromolecule #2: Neogenin
+Macromolecule #3: Repulsive Guidance Molecule B (C-terminal region)
+Macromolecule #4: RGM domain family member B
+Macromolecule #5: CALCIUM ION
+Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.07 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 10 mM HEPES pH 7.5, 150 mM NaCl, 2 mM CaCl2, 1 mM sucrose octasulfate, 0.01% NaN3 | ||||||||||||||||||
| Grid | Model: PELCO Ultrathin Carbon with Lacey Carbon / Material: COPPER / Support film - Material: CARBON / Support film - topology: LACEY / Support film - Film thickness: 3 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER / Pretreatment - Pressure: 101.325 kPa Details: Agar Scientific Ultra-thin carbon support film, 3 nm - on lacey carbon. https://www.agarscientific.com/ultra-thin-carbon-support-film-3-nm-on-holey-carbon | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: Lacey carbon grids with 3 nm ultrathin carbon support film were glow discharged for 30 seconds at high RF level using Harrick Plasma Cleaner, model PDC-002-CE, and then 3.5 microl of the ...Details: Lacey carbon grids with 3 nm ultrathin carbon support film were glow discharged for 30 seconds at high RF level using Harrick Plasma Cleaner, model PDC-002-CE, and then 3.5 microl of the sample was pipetted per grid. Excess protein was blotted away for 3 seconds using filter paper (round filter paper for Vitrobot from Agar Scientific, catalogue number 47000-100) and Vitrobot Mark IV (Thermo Fisher Scientific) (relative force -15) at 95-100% humidity. Grids were plunge frozen in liquid ethane.. | ||||||||||||||||||
| Details | The ternary NEO1-NET1-RGMB complex was purified by SEC on a S200 10/300 Increase column with a running buffer of 10 mM HEPES pH 7.5, 150 mM NaCl, 2 mM CaCl2, 1 mM sucrose octasulfate, 0.01% NaN3 at 4 degrees C . The peak fraction containing the ternary complex was diluted to 0.07 mg per ml in SEC buffer. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 77.0 K |
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1635 / Average exposure time: 37.26 sec. / Average electron dose: 40.0 e/Å2 Details: Cryo-EM data were collected on a Titan Krios G3i microscope (Thermo Fisher Scientific) operating at 300 kV with a 50 microm C2 aperture and Volta phase plate (Thermo Fisher Scientific), at ...Details: Cryo-EM data were collected on a Titan Krios G3i microscope (Thermo Fisher Scientific) operating at 300 kV with a 50 microm C2 aperture and Volta phase plate (Thermo Fisher Scientific), at the Division of Structural Biology, University of Oxford. Movies were recorded using a FEI Falcon III direct electron detector in electron counting mode using EPU software at a nominal magnification of 96000x, corresponding to a physical pixel size of 0.85 angstrom/pixel. A total dose of 40 electrons per square angstrom was used at a dose rate of 0.77 electrons/pix/sec. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: -0.7000000000000001 µm / Nominal defocus min: -0.5 µm / Nominal magnification: 96000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Crystal structure used as a model for fitting and rigid body refinement will be described by Robinson, Griffiths, van de Haar, Malinauskas et al., Cell, 2021. |
|---|---|
| Refinement | Protocol: RIGID BODY FIT / Target criteria: Correlation coefficient |
| Output model |  PDB-7ndg: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)